Preliminary Characterization of Xylose Reductase Partially Purified by Reversed Micelles from Candida tropicalis IEC5-ITV, an Indigenous Xylitol-Producing Strain ()
1. Introduction
Extraction by reverse micelles is a useful versatile tool for separating biomolecules such as enzymes. This method involves the use of ternary mixtures composed of water-surfactant-solvent which in a first step (forward extraction), through agitation, leads to the aggregation of surfactant molecules that form a hydrophilic core (reverse micelles) where the enzyme to be purified can be integrated. In a second step (backward extraction), through its transfer to a new aqueous phase the enzyme of the reverse micelles is released, so that a quantitative recovery of purified protein is observed [1]. The extraction of enzymes by reverse micelles has been demonstrated for a number of enzymes in various reverse micelles systems: porcine pepsin and bovine chymosin [2], alcohol dehydrogenase [3], xylanase [4], β xylosidase [5], glucose oxidase [6], nattokinase [7], xylitol dehydrogenase [8,9] and xylose reductase [9-11].
Yeast aldose reductases belong to the aldo-keto reductase superfamily of enzymes whose members are responsible for a wide variety of biological functions in organisms that contain them. Thus, xylose reductase (EC 1.1.1.21) catalyzes the reversible reduction of D-xylose to xylitol in the cytoplasm of the yeast that consumes or ferments pentoses. The xylitol thus produced can be oxidized to xylulose or released into the environment depending on organism culture conditions [12].
Xylitol is a valuable sweetener which has applications in the food and pharmaceutical industries. Its biotechnological production from yeast and agro-industrial waste hydrolysates, such as sugar cane bagasse, rich in D-xylose, are a cheaper alternative production of this sweetener, which is currently synthesized by chemical methods [13]. Xylose reductase is one of the key enzymes for xylitol production, so understanding the mechanisms that regulate its activity could aid in establishing optimal conditions for xylitol production. This research evaluates the effectiveness of reversed micelles in the extraction and purification of xylose reductase from the xylitol producing indigenous yeast Candida tropicalis IEC5-ITV grown in hydrolyzed sugar cane bagasse-like synthetic medium, in addition to determining some biochemical characteristics of this enzyme.
2. Materials and Methods
2.1. Microorganism and Inoculum Preparation
Candida tropicalis IEC5-ITV was isolated in the Bioengineering Laboratory of the Veracruz Institute of Technology. The yeast was maintained at 4˚C on agar (25.0 g/L), yeast extract (10.0 g/L) and xylose (20.0 g/L). The inoculum was grown by transferring three loops of the organism to a 500 mL Erlenmeyer flask with 250 mL synthetic medium (xylose 20.0 g/L, KH2PO4 5.0 g/L, (NH4)2SO4 2.0 g/L, Mg(SO4) 7H2O 0.4 g/L and yeast extract 1.0 g/L), pH 5.5 and incubated with agitation at 30˚C, 24 h, 250 rpm [14].
2.2. Fermentation Conditions and Crude Extract Obtention
A 500 mL Erlenmeyer flask containing 300 mL hydrolyzed sugar cane bagasse-like synthetic medium xylose (30 g/L), glucose (2.68 g/L), arabinose (3.4 g/L), acetic acid (5.0 g/L) and furfural (<0.05 g/L), was inoculated with 6 × 106 viable cells/mL. This medium also enriched with urea (3.0 g/L), KH2PO4 (5.0 g/L) and yeast extract (1.0 g/L), pH adjusted to 5.5 was then incubated with agitation at 30˚C, 24 h, 250 rpm [14]. The cells were harvested by centrifugation at 2700 g (Hermle Z 300 K®) for 20 min at 4˚C. The biomass was washed with deionized water, weighed and resuspended in 0.1 M phosphate buffer, pH 7, 40% alumina (w/w) was added and stored at –70˚C, then thawed and macerated with mortar and pestle until at least 85% broken cells were observed under the microscope in a Thoma chamber. The cell homogenate was centrifuged at 2700 g for 20 minutes at 4˚C. The supernatant (crude extract) was used for subsequent enzyme assays. The protein concentration was determined by the Bradford method, using bovine serum albumin as a standard [15].
2.3. Extraction and Purification of Xylose Reductase (XR) in Reverse Micelles
Extraction and purification was performed according to Cortez et al. [11], except that electrical conductivity was not adjusted. Three mL crude extract (CE) was mixed with an equal volume of micellar microemulsion, composed of cetyl trimethyl ammonium bromide (CTAB)/ isooctane 0.15 M, hexanol 5%, 22% butanol, pH 7 to 5˚C. This mixture was mechanically stirred (vortex mixer) for 1 min to obtain equilibrium then centrifuged at 1735 g for 10 min at 5˚C. Two phases were obtained (micellar phase I and aqueous phase I). To transfer the enzyme contained in the micellar phase I to a fresh aqueous phase, 2 mL micellar solution were mixed with 2 mL 1.0 M acetate buffer, pH 5.5, 1.0 M NaCl, stirred with a vortex mixer for 1 min, then centrifuged at 1735 g for 10 min at 5˚C to separate the aqueous phase II (APII) from the micellar solution. Two clear phases were obtained, with a small amount of white material collected at the interface. Determinations were made for protein concentration and enzyme activity for both aqueous phases I, II and CE.
2.4. Enzyme Activity
Reaction medium constituents for the measurement of xylose reductase (XR) activity were: deionized water 100 μL, 0.1 M phosphate buffer, pH 7.0, 600 μL, 0.1 M β- mercaptoethanol, 100 μL, 50 μL enzyme extract, 50 μL 3.0 M NADPH. The reaction was initiated with the addition of 100 μL 0.5M D-xylose, and NADPH consumption was followed by the change in absorbance for 1 minute at 340 nm (Genesys 10 UV-VIS, Thermo Scientific) [16]. Activity determination was performed in duplicate on at least 2 crude extracts obtained at different times. One unit (U) XR activity is defined as the amount of enzyme required to catalyze the formation of 1 µmol NADP+ per min using an extinction coefficient of 6.22 × 10–3 M–1·cm–1. Each experiment was performed in triplicate, and the results were expressed as average values.
2.5. Effect of pH and Temperature on Enzyme Activity and Stability
XR activity was evaluated at room temperature by varying the pH of the enzyme assay using appropriate 0.1 M buffer from 4.5 to 8.5. Stability was determined by incubating the enzyme solution 1 h at room temperature to the desired pH before the standard test to determine XR activity. The temperature of the enzymatic assay varied from 30˚C to 60˚C. Enzyme stability was evaluated by incubating it for 1 h at the desired temperature before activity determination [17].
2.6. Determination of KM and VMax
Apparent XR kinetic constants were determined by the Lineweaver-Burke method, changing substrate and cofactor concentrations at intervals from 0.0 M to 0.25 M for xylose, and 0.03 mM to 0.18 mM for NADPH according to the standard assay [17].
2.7. Enzyme Stability during Storage
CE and purified enzyme (PAII) samples were stored at –18˚C. After 2 months, enzyme activity was determined in both samples at room temperature.
2.8. Molecular Weight Determination of Xylose Reductase
The protein present in the crude extract and the aqueous phase II were concentrated by ultrafiltration at 4˚C using a stirred Amicon cell 8200 equipped with a PM 10 filter (cut off 10,000). Sodium dodecyl sulfate-polyacrylamide gel elelectrophoresis (SDS-PAGE) [15] was performed using a Bio-Rad Mini Protein electrophoresis unit, using a molecular weight marker (Bio-rad® Precision Plus Dual Color Standards ProteinTM). A 0.025 mL amount of a 2 mg/mL protein solution was loaded per well. The gel was stained with Coomassie Brilliant Blue R 25.
3. Results and Discussion
3.1. Extraction and Purification of Xylose Reductase by Reverse Micelles
The extraction process by reverse micelles is mainly determined by electrostatic interactions between a charged protein and the micellar wall. Protein transfer during the forward extraction only takes place when aqueous phase pH is such that the net charge on the surface of the protein is electrically opposite to that of surfactant [6]. The isoelectric point (pI) of XR produced by Candida tropicalis IEC5-ITV is unknown, although a pI between 4.10 and 4.15 has been reported for Candida tropicalis [16]. Therefore, the XR described in the present study, at the pH used in the forward extraction (pH 7), can have a negative global charge, and the surfactant, CTAB, under these conditions, a positive charge which favors XR inclusion in the reverse micelles. In the backward extraction, the pH value must allow the protein to have the same charge as the surfactant molecules; in our case, the pH of the backward extraction was 5.5 which favored the positive charge of XR. When the surfactant has the same charge, protein repulsion forces are created and the micellar diameter is decreased, causing the release of the protein [9]. Electrostatic interactions, which may cause enzyme migration to the micellar core, are one of the most prevalent factors in reverse micelles extraction and explain the high recovery of XR in this study (Table 1). XR recovery was over 100% since all the activity of the enzyme present in the crude extract was transferred to the fresh aqueous phase after backward extraction, no activity of XR was detected in API. A range of transfer efficiencies have been found including 100% efficiency or complete transfer into reverse micelles [2].
Using reversed micelles of BDBAC [N-benzyl-N-dodecyl-N-bis (2-hydroxyethyl) ammonium chloride] cationic surfactant, Cortez et al. [10] recovered about 136% XR from C. guilliermondii FTI 20037 and the same researchers using CTAB [9-11] and the same organism recovered 100% XR. A purification factor of 8.1 fold obtained in this study compares favorably to those mentioned above (4.1 and 5.6, respectively). This difference can be attributed to variations between protocols. The main differences were fermentation medium (a hydrolyzate of sugarcane bagasse concentrate containing 42 g·L–1 xylose and controlled aeration conditions versus a synthetic medium with 30 g·L–1 xylose and uncontrolled air flow) and adjustment of electrical conductivity of the micellar microemulsion (carried out or not). The interacttion of the surfactant with any broth constituents may affect the preferential separation of the desired product [7] and many publications have demonstrated that different ionic strength affects the ability of reverse micelles to entrap the protein [1].
3.2. Effect of pH on Enzyme Activity and Stability
A pH value about of 6.0 was the optimum pH for XR activity before extraction (Figure 1).
This result is in agreement with those reported in the literature for other XRs, insofar as the optimum pH for activity was 6.0 for P. tannophylus [18], P. stipitis [19] and Candida tropicalis [16-20] and 5.5 and 6.0 for Candida guilliermondii FTI 20037 [17].

Table 1. Purification of XR produced by C. tropicalis IEC5- ITV using reverse micelles.
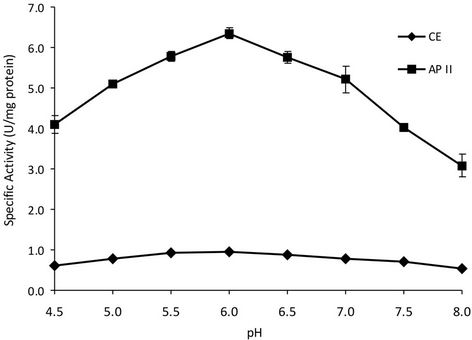
Figure 1. Effect of pH on xylose reductase activity present in crude extract (CE) and purified by reverse micelles (AP II).
The XR of Candida tropicalis IEC5-ITV in AP II also exhibited a peak of maximum activity at pH 6.0; Cortez et al. [11] found that the pH optimum for XR activity for Candida guilliermondii FTI 20037 before and after extraction with reversed micelles was about 6.0, however maximum XR activity diminished about 44% after submission to the extraction compared to that found in crude extract, attributing this to the effect of the solvents used in this method, which may affect the conformation of the protein. This was not the case in this study.
The residual specific activity (Table 2) of the XR of Candida tropicalis IEC5-ITV after one hour of incubation at different pH values, reached an optimum of 7 before and after extraction with reverse micelles, the decrease in activity being attributed to damage in enzyme structure.
3.3. Effect of Temperature on Enzyme Activity and Stability
On the other hand, the specific activity of XR from Candida tropicalis IEC5-ITV in the crude extract has an optimum at 40˚C, while the enzyme purified by reverse micelles reaches a temperature of 30˚C, decreasing by 38.5% at 40˚C (data not shown). The effect of temperature on the stability of XR of Candida tropicalis IEC5- ITV was similar in CE and AP II, since in both fractions enzyme activity is maintained above 90% between 30˚C - 40˚C, however after incubation for one hour at 50˚C lost residual specific activity retains 16.6% and 17.85% respectively (Table 3). This decline in enzyme activity due to temperature is attributed to enzyme denaturation and/or, in the case of crude extract, to activation of those proteases present. Similar results were found for XR of C. guilliermondii FTI 20037 obtained by reverse micelles [11].
3.4. Determination of KM and VMax
The values obtained for the kinetic parameters Michaelis-Menten constant (KM) and maximum velocity reaction (VMax) for XR of C. tropicalis IEC5-ITV, showed the following characteristics: the KM for xylose increased 77.3% after extraction (0.0059 M to 0.026 M), while the KM for NADPH was 92.9% lower (12.04 mM to 1.85 mM). The same applies to the VMax for xylose (0.42 U/L in CE and 4.53 U/L in AP II). The same was found by Cortez et al. [11] for XR of C. guilliermondii purified by reverse micelles, only to a lesser extent. In this work the kinetic constants were significantly affected. This can be explained, at least in part, because the enzyme molecules are disrupted in structure during the extraction process, so that as the affinity for the substrate decreases, that for the coenzyme increases [11]. Activity of XR for NADH was not detectable, as reported for XR of C. tropicalis IFO 0618 [16].
3.5. Enzyme Stability during Storage
After two months’storage at –18˚C, enzyme activity before purification remained intact and the purified enzyme lost 76.60% activity (Figure 2), probably due to its instability given by the contact with solvents during extraction.
3.6. Molecular Weight Determination of Xylose Reductase
Enzyme analysis by SDS-PAGE (Figure 3) showed a band present in AP II which coincided in mobility with one present in CE, both with a molecular weight of 32.42

Table 2. Effect of pH on stability of the XR of Candida tropicalis IEC5-ITV.
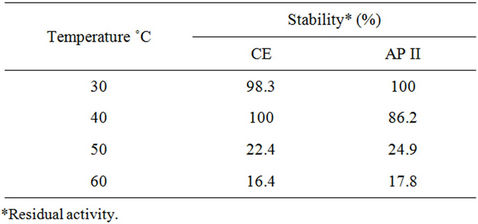
Table 3. Effect of temperature on stability of XR of Candida tropicalis IEC5-ITV.

Figure 2. Residual activity of xylose reductase present in the crude extract (CE) and the aqueous phase II (AP II) of Candida tropicalis IEC5-ITV after two months of storage.

Figure 3. SDS-PAGE of C. tropicalis. Lane 1: marker proteins. Lane 2: micellar extract (AP II). Lane 3: crude extract (CE).
kD which is within the molecular weight range reported for other XRs [20-22]. XRs have been found to be present also as a dimer with subunits approximately 36 kD in molecular weight [20-23] so it is necessary in our case to perform electrophoresis under native conditions to determine whether the XR of Candida tropicalis IEC5- ITV is composed of one or more subunits. The presence of two additional bands, with molecular weights of 54.95 and 66.06 kD, was also noted but to a lesser extent, indicating that XR is not the only protein extracted under these conditions. This suggests that these two proteins have the size and appropriate charge to be included in the reverse micelle. Hence, a further step of purification is required to obtain a homogeneous product.
4. Conclusions
The use of reversed micelles for the extraction and purification of xylose reductase from Candida tropicalis IEC5-ITV resulted in a considerable saving of time and reagents. The recovery yield obtained in this work was 100% and the enrichment factor was 8.1. A partial preliminary characterization of the enzyme was achieved, showing that the kinetic parameters are modified when the enzyme is purified, the KM increasing for xylose and decreasing for NADPH. A further step of purification and structure sequencing will allow us to better understand their stability and kinetic changes observed in this study.
5. Acknowledgements
This work is part of the project entitled “Technology for sustainable development in the production of xylitol and energy from sugarcane bagasse”, No. 31346. Fondo Mixto CONACYT-Gobierno del Estado de Veracruz. (National Council of Science and Technology-Veracruz State Government Mixed Fund). Authors acknowledge the critical reading of Patricia Hayward Jones MSc. and Dulce María Barradas Dermitz MSc.
NOTES