Iterated Function System-Based Crossover Operation for Real-Coded Genetic Algorithm ()
1. Introduction
Genetic algorithm (GA) [1] is a stochastic search algorithm for solving optimization problems. It can help find out the globally optimal solution over a domain. In general, three genetic operations affect the performance of the GA: selection, crossover and mutation. Selection operation selects the parents from the population with respect to some probability distribution and the fitness values. The crossover operation combines the information of the selected parents and generates the offspring. The mutation operation introduces changes to the offspring variables. Recently, different crossover operations for real-coded GA have been proposed to improve the efficiency. The unimodal normal distribution crossover (UNDX) was proposed [2] for multi-modal and highly epistatic functions. The blend crossover (BLX-α) [3] was reported showing good search ability for separable fitness functions. Average-bound crossover [4] was introduced to enhance the solution quality and solution stability. In this paper, a new crossover operation is presented.
Iterated function system (IFS) theory was proposed by Barnsley [5] , which involved a specific fractal that enhances a self-similarity property. Based on the IFS, objects are dissected into pieces that are similar to the whole object. Taking advantage of the self-similarity property of IFS, an iterated function system crossover (IFSX) is proposed for real-coded GAs. It will be shown that the GA with IFSX will perform more efficiently and provide a faster convergence in a suite of benchmark test functions.
2. Iterated Functions System Crossover
The idea of using IFSX to reproduce offspring is shown in Figure 1. The procedure is as follows:
1) Select 2 parents 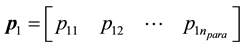 and
and 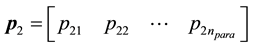 from the population, where
from the population, where 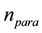 is the number of parameters.
is the number of parameters.
2) Combine the information (genes) of 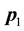 and
and  to form a vector v of complex elements given by
to form a vector v of complex elements given by
 (1)
(1)
3) Based on the IFS theory [4] , let
 (2)
(2)
where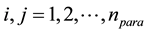 ,
, 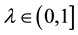 is a scaling factor.
is a scaling factor. 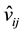 are the possible values generated by the IFS that
are the possible values generated by the IFS that
exhibits a self-similarity property. For instance, in Figure 1, there are 3 values, v1, v2 and v3. From (2), we have
9 values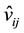 , i, j = 1, 2, 3. This figure shows a fractal, and the patterns of
, i, j = 1, 2, 3. This figure shows a fractal, and the patterns of  and
and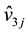 , j = 1, 2, 3 are similar to the pattern of v1, v2 and v3. In some cases, the value of
, j = 1, 2, 3 are similar to the pattern of v1, v2 and v3. In some cases, the value of 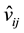 may be out of the boundary. Then, the system
may be out of the boundary. Then, the system
will generate a random value (within the boundary) to replace it.
4) Randomly pick up npara variables from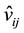 . If
. If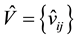 ,
,
then ![]() (3)
(3)
For example, in Figure 1, a possible Q can be![]() .
.
5) Generate the offspring ![]() and
and ![]() as follows:
as follows:
![]() (4)
(4)
![]() (5)
(5)
where Re(Q) and Im(Q) generate vectors formed by the real part and imaginary part of the elements of Q respectively.
Crossover operation is mainly for exchanging information from the two selected parents. In traditional crossover operations (e.g. UNDX and BLX-α), the information is exchanging gene by gene. In the proposed IFSX, each offspring gene is affected by all other genes of the parents, which is a more “complete” crossover operation for information exchange. The IFSX crossover makes the GA operation performs better in terms of fitness value and convergence rate.
3. Simulation Results
The GA with the proposed IFSX goes through six test functions. The results are compared to those from GAs with UNDX and BLX-α. For each test function, the population size is 50 and all the simulation results are averaged ones out of 50 runs. The selection algorithm and the mutation operation are the roulette wheel selection [1] and the non-uniform mutation [1] respectively. The six test functions are listed in Table 1. f1 is a sphere model which is smooth and symmetric. f2 is a step function, which is a representative of flat surfaces. f3 is a quartic function which is a simple unimodal function padded with noise. f4 is a Shekel’s foxholes function and f5 is a Kowalik’s function, which are multimodel functions with only a few local minima. f6 is an Ackley’s function which is a multimodel function with many local minima. The parameter λ of the IFSX are set at 0.005, 0.001, 0.001, 0.01, 0.005, 0.005 for f1 to f6 respectively, and the parameter of the BLX-α is set at 0.336 [3] . The simulation results obtained by the GA with the proposed IFSX, UNDX and BLX-α are shown in Figure 2 and Table 2. It can be seen that the searching performance of the proposed IFSX is improved with faster convergence rate.
![]()
Table 1. Six benchmark test functions.
![]()
Table 2. Statistical results for f1 to f6.
4. Conclusion
In this paper, a new crossover of IFSX for real-coded GA has been proposed. Take the advantage of the iterated function system theory and integrate into crossover operation of real-code genetic algorithm, the solution quality of the searching is enhanced. A suite of benchmark test functions has been used to illustrate the merits of the IFSX.