Molecular Characterization and mRNA Expression of Cytochrome P450 1A1 and Cytochrome P450 3A in Liver of Kafue Lechwe (Kobus leche kafuensis) as Potential Biomarkers of Pollution of the Kafue River Basin, Zambia ()
1. Introduction
The Kafue River basin in Zambia is one of Africa’s most diverse ecological habitats. It comprises Lochnivar National Park in the South, Blue Lagoon National Park in the North and Kafue Basin Game Management Areas (GMAs) [1]. It is home to over 400 species of birds, a variety of fish species and it provides for extensive endemism of rare mammals such as the Kafue lechwe (Kobus leche kafuensis) antelope [1]. In fact, the Kafue lechwe is unique to this habitat and is naturally found nowhere else in the world. It is one of the most hunted species for game meat consumption and therefore has an impact on the health of the local communities and the nation at large. As a result, the Kafue river ecosystem constitutes a unique environment of social, economic and environmental importance, and international acclaim and recognition [2]. Unfortunately, because of the location of Kafue river basin, a lot of effluents from industries, mining and agricultural activities have been documented thus exposing the basin to potential destruction. The Kafue River has been reported to have higher than normal levels of pollutants compared to the average world river [3]. Pettersson and Ingri (2000) [4] and Pettersson et al. (2000) [5] demonstrated the increased concentrations of dissolved and suspended heavy metals in the Kafue River and the marked accumulation of cobalt, copper, iron and manganese within the river sediment. Norrgren et al. (2000) [6] reported a high concentration of several pesticides, i.e. dichlorodiphenyltrichloroethane (DDT) with its major metabolites, polychlorinated biphenyl (PCB), and dieldrin in Kafue river water. Syakalima et al. (2006) [7] reported a widespread contamination of the Kafue river with pesticides/herbicides such as DDT, chlordane, toxaphene, terbufos, kelthane, endosulfan, dieldrin, 1-1 dichloro-2,2 bis (p-chlorophenyl) ethane (pp’-DDD), atrazine, and endrin. Sichilongo and Torto, (2006) [8] also demonstrated tissue contamination of the Kafue lechwe by endocrine disruptors such as deltamethrin, heptachlor, aldrin, d-t-allethrin, endosulfan, Dichlorodiphenyldichloroethylene (pp-DDE), dieldrin, endrin, pp’-DDD and DDT in the Lochinvar National Park. Despite pollutants being well documented, there have been few studies to demonstrate the actual biological effect on wild animals in the Kafue river basin. The cytochrome P450 enzymes (CYPs) have been widely used in mammals to demonstrate biological effect of pollutants since they play a key role in detoxification of many chemicals [9-11]. Of the numerous CYP families identified, Cytochrome P450 1A1 (CYPlAl) has been studied in many species of different taxa and its induction mediated by the aryl hydrocarbon receptor [12] is widely used as a biomarker of exposure to polycyclic aromatic hydrocarbons (PAH), PCB and polychlorinated dibenzop-dioxin (PCDD) [13]. The Cytochrome P450 3A (CYP3A) subfamily has mostly been investigated in mammals [14] and is involved in the metabolism of environmental pollutants such as DDT and its metabolites and as well as therapeutic drugs [15]. The CYP1A1 and CYP3A are therefore important indicators of environmental pollution and fundamental enzymes for detoxification/activation of xenobiotics. Their gene expression levels have been used extensively worldwide in biomonitoring programs for detecting exposure to environmental pollutants but have rarely used in Zambia. Their applicability in Kafue lechwe is still not done. The aim of this study was to characterize CYP1A1 and CYP3A in liver of the Kafue lechwe from Lochnivar and Blue Lagoon GMAs and to determine their mRNA expression levels as potential biomarkers of pollution. Furthermore, the study determined their molecular phylogeny covering CYP1A1 and CYP3A genes of other animals.
2. Methods
2.1. Samples from Kafue Lechwe
Twenty three and fifteen Kafue lechwe liver samples were collected from Lochnivar and Blue Lagoon GMAs respectively in 2009 (Figure 1). The Zambia Wildlife Authority issued the permit to obtain samples from Kafue lechwe for disease surveillance. Postmortems of the animals were carried out as described by Gracey and Collins (1992) [16]. Dissections of the liver samples were performed and put in liquid nitrogen. The samples were then stored at −80˚C in the laboratory after transportation from the field.
2.2. RNA Isolation and cDNA Synthesis of CYP1A1 and CYP3A
The preparation of total RNA from individual liver samples was done by the single-step method [17] using TRI reagent (Sigma Chemical Co. St. Louis, MO, USA). Spectrophotometric method using Nanodrop ND-1000 was used to determine the concentration and purity of RNA at 260 and 280 nm [18]. Synthesis of cDNA from total RNA was carried out by reverse transcription as
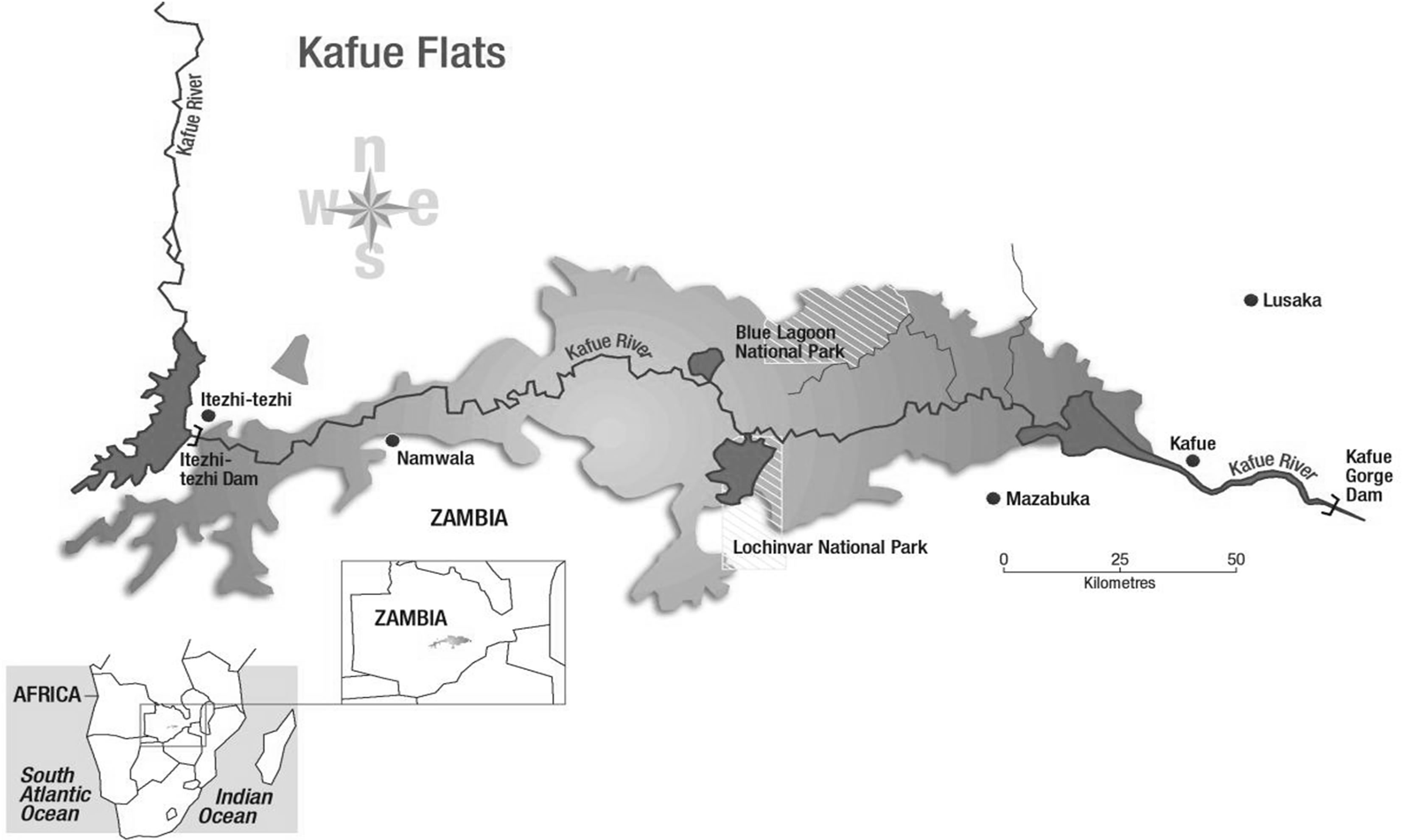
Figure 1. Lochnivar and blue lagoon game management areas.
described in literature [19].
PCR was performed on Bio-Rad icyclerTM Thermal Cycler. In the amplification, the primers (Sigma Genosys, Hokkaido, Japan) which were selected to amplify 252 bp fragments of lechwe CYP1A1 cDNAs were designed by first searching the CYP1A1 sequence of horse CYP1A1 (accession number XM_001493909), pig CYP1A1 (accession number NM_214412), and sheep CYP1A1 (accession number NM_001129905) from NCBI. Clustal W [20] was used to align and determine the common (conserved) sequence of the three species. The conserved sequences were used as primers because it was thought that they were similar to Kafue lechwe CYP1A1 sequence and the following primers were designed and used: Forward >5’-ctttgtgaaccagtggcaga-3’ and Reverse >5’-ggccaggaagagaaagacct-3’. The primers used to amplify 500bp of CYP3A cDNAs were CYP 3A mammalian degenerate primers: Forward (5’-tttggrgcctacagcatgga-3’ r = a or g) and Reverse (5’-acyaccatgtcragatactcc-3’ r = a or g, and y = t or c) were both positioned upstream from the well described cysteine-containing heme-binding region [21-24]. The PCR reaction was prepared and thermalcycler conditions set as described in literature [19] but with an annealing temperature of 57˚C. Electrophoresis of PCR products was run on 1.5% agarose gel stained with ethidium bromide (0.25 μg/ml) in TBE buffer and single bands of 252 bp and 500 bp were separated for CPY1A1 and CYP3A respectively.
2.3. Sequencing of CYP1A1 and CYP3A
Direct sequencing of CYP1A1 and CYP3A was performed using PCR products (60 to 100 ng/μl) and Big Dye Terminator kit according to the manufacturer’s instructions (Applied Biosystems, CA, USA). After amplification, ethanol precipitation was performed and the sequences were analyzed by the automated DNA sequencer (ABI Prism 310 genetic analyzer, Applied Biosystems). BLAST searching against the NCBI sequence database was utilized to confirm sequence identities.
2.4. Phylogenetic Relationships of CYP1A1 and CYP3A
Phylogenetic relationships were calculated by Clustal W [20]. Kimura’s 2-parameter model [25] was used to construct the phylogenetic tree for nucleotide sequences of lechwe CYP1A1 by the NJ method and was based on fragments of CYP1A1 of other animal species. The following nucleotide sequences and their GeneBank accession numbers were used for phylogeny; Ovis aries (Sheep) Cytochrome P450 1A1 NM_001129905, Bos taurus (Cattle) Cytochrome P450, family 1, subfamily A (aromatic compound inducible), polypeptide 1 (CYP1A1), mRNA XM_588298, Balaenoptera acutorostrata (Minke whale) CYP1A1 AB_231891, Lagenorhynchus acutus (Atlantic white-sided dolphin) CYP1A1 AY641536, Stenella coeruleoalba (Striped dolphin) CYP1A1 AF235141, Sus scrofa (Pig) Cytochrome P450 1A1, (CYP1A1) mRNA NM_214412, Danio rerio (Zebrafish) cytochrome P450 1A1 (CYP1A1) mRNA AF210727, Oryzias latipes (Medaka) cytochrome P450 1A (CYP1A) mRNA AY297923. Phylogenetic tree for CYP3A deduced amino acid sequences was constructed by the Maximum likelihood method based on MEGA 5 program [26]. The following sequences and their GeneBank accession numbers were used in the phylogeny; Ovis aries (Sheep) Cytochrome P450 subfamily 3A, polypeptide 24 (CYP3A24) NM_001129904, Bos taurus (Cattle) CYP3A24 XM_002698141, Bos taurus (Cattle) Cytochrome P450, subfamily 3A, polypeptide 4 (CYP3A4) NM_001099367, Bos taurus (Cattle) CYP3A4-like XM_002703144, Bos taurus (Cattle) CYP3A5 NM_001075888, Balaenoptera acutorostrata (Minke whale) CYP3A32, AB014349, Balaenoptera acutorostrata (Minke whale) CYP3A72 AB290009, Sus scrofa (Pig) CYP3A22 NM_001195509, Sus scrofa (Pig) Cytochrome P450 3A29, (CYP3A29) mRNA NM_214423, Sus scrofa (Pig) CYP3A39 NM_214422, Sus scrofa (Pig) CYP3A46 NM_001134824, Canis lupus familiaris (Dog) CYP3A12 XM_846627, Canis lupus familiaris (Dog) CYP3A26 NM_001003338, Canis lupus familiaris (Dog) LOC479740 XM_536868, Canis lupus familiaris (Dog) LOC489851 XM_546969, Equus caballus (Horse) CYP3A93 NM_001190938, Equus caballus (Horse) Cytochrome P450 3A94 (CYP 3A94), mRNA NM_001101651, Equus caballus (Horse) CYP3A95 NM_001190940, Equus caballus (Horse) CYP3A96 NM_001146163, Equus caballus (Horse) CYP3A97 FN669297, Homo sapiens (Human) CYP3A4 DQ924960, Homo sapiens (Human) CYP3A5 NM_000777. The number of bootstrap replications was set to 500. Tamura-Nei model was applied in nucleotide sequences and JTT model was applied in amino acid sequences. PAML [27] was used to test the differences in selective pressure among Bovidae CYP3A genes. The dN/dS ratio of other nuclear gene, MC1R, was also estimated. A phylogenetic tree based on genomic sequence for user tree was used. The F3x4 codon model was applied and allowed for estimating synonymous and nonsynonymous substitution (dN/dS) rate.
2.5. Quantification of CYP1A1 and CYP3A mRNAs
Quantitative comparative (relative) real-time RT-PCR was used to measure the expression levels of CYP1A1 and CYP3A mRNAs. Two microliter of 25 fold cDNA and DyNamoTM HS SYBR Green qPCR Kit were used according to the manufacturer’s instructions (Finnzymes, Espoo, Finland). PCR amplification and data analysis (Step one software version 2.0) was performed on a step one plus real-time PCR system (Applied Biosystems, Japan). A set of specific primers (Sigma Genosys) for Kafue lechwe CYP1A1 and CYP3A were designed using software ‘‘primer 3” (http://frodo.wi.mit.edu/primer3/) using the lechwe CYP1A1 and CYP3A sequences obtained. This resulted in several candidates of primer sets which were experimentally tried (data not shown) and the following primer sets were used in the experiment: CYP1A1-Forward >5’-caatggtctcaccaatgcac-3’, CYP1A1- Reverses >5’-tgaaccagtggcagatcaac-3’, CYP3A-Forward >5’-ccacgtgtggactttcttca-3’, CYP3A-Reverse >5’-tggtctcatagccagcaaaa-3’. The cDNAs were amplified with GAPDH (accession number BC102589) [28] as a control housekeeping gene. The Forward primer was (5’-ggcgtgaaccacgagaagtataa-3’) and the reverse primer (5’-ccctccacgatgccaaact-3’) (Sigma Genosys, Hokkaido, Japan). Conditions for quantitative real-time (RT)-PCR were as described in literature [19]. Electrophoresis was run on 1.5% agarose gel stained with ethidium bromide (0.25 μg/ml) in TBE buffer and single bands of 167 bp, 130 bp and 120 bp were separated for CYP1A1, CYP3A and GAPDH, respectively.
2.6. Statistical Analysis
The data obtained were normalized by base 10 logarithm transformations and regional differences in CYP1A1 and CYP3A mRNA expressions were analyzed by Student’s t test (p < 0.05). Statistics analysis of data was conducted using JMP 9 (SAS Institute, Cary, NC, USA).
3. Results
3.1. Characterization of CYP1A1 and CYP3A in Kafue Lechwe
The partial nucleotide sequences of lechwe CYP1A1 and CYP3A were 206 bp and 383 bp respectively (Figure 2). The sequences were deposited in GenBank and were assigned the following accession numbers; AB742159 and AB742160 for lechwe CYP1A1 and CYP3A respectively.
3.2. Phylogenetic Relationships of Kafue Lechwe CYP1A1 and CYP3A
Lechwe CYP1A1 showed higher identity with sheep_ ORF (98%), then cattle_ORF (97%), followed by that of minke whale_ORF (91%) and lastly dolphins_ORF (87%). The deduced partial nucleotide sequence of Kafue lechwe CYP3A revealed higher identities with sheep ORF (97%), then cattle _ORF (95%), but shared lower identities with horse _ORF (84%) and pig _ORF (83%). Phylogenetic tree constructed from Cetruminantia CYP1A1 nucleotide sequences (Figure 3) revealed that Kafue lechwe CYP1A1 was clustered with that of sheep and cattle CYP1A1. The CYP1A1 phylogenetic tree was corresponding to Cetruminantia phylogeny. The phylogenetic tree of mammalian CYP3A was constructed from deduced amino acid sequences and revealed that CYP3A of Kafue lechwe was clustered with that of sheep CYP3A24 and cattle CYP3A24 and other CYP3As were clustered in each species (Figure 4).
The dN/dS ratio of Bovidae CYP3As was estimated by PAML. In Bovidae CYP3A cluster there was no branch that was estimated to be significantly higher than one. The ratio of lechwe CYP3A was estimated at 0.67 (Figure 5). The dN/dS ratio of other nuclear genes was also estimated for comparison. The dN/dS ratio of lechwe MCR1 was 0.02.
3.3. Quantities of Kafue Lechwe CYP1A1 and CYP3A mRNAs in Lochnivar and Blue Lagoon GMAs
The mean and standard error about the mean for mRNA expression levels of CYP1A1 and CYP3A in lechwe from Lochnivar GMA (n = 23) were 7.64 ± 2.85 and 1.27 ± 0.50 respectively. In Blue Lagoon GMA (n = 15) the mean and standard error about the mean for mRNA levels of CYP1A1 and CYP3A were 2.67 ± 1.37 and 5.37 ± 1.62 respectively. There was a 2.9-fold increase in CYP1A1 mRNA expression level in Kafue lechwe from Lochnivar GMA when compared with that of Kafue lechwe from Blue Lagoon GMA.
The difference was significant (P < 0.05) (Figure 6). There was a 4.2-fold increase in CYP3A mRNA expression level in Kafue lechwe from Blue Lagoon GMA when compared with that of Kafue lechwe from Lochnivar GMA and the difference was significant (P < 0.05) (Figure 6).
4. Discussion and Conclusions
Studies carried out in the Kafue River have established the presence of several classes of pollutants including heavy metals [5] and pesticides [6]. Accumulation of these contaminants in animals such as the Kafue lechwe that are dependent on the Kafue River has also been reported [7,8]. The presence of such contaminants in the animals can cause known adverse effects that include reproductive and immunological abnormalities as well as numerous pathologies [29]. Unfortunately, no studies have been conducted to demonstrate similar effects in the Kafue lechwe. Knowledge of xenobiotic metabolism in these animals is necessary to precisely evaluate their susceptibility to effects of these chemicals after their

Figure 2. Partial nucleotide sequences of Kafue lechwe CYP1A1 and CYP3A.
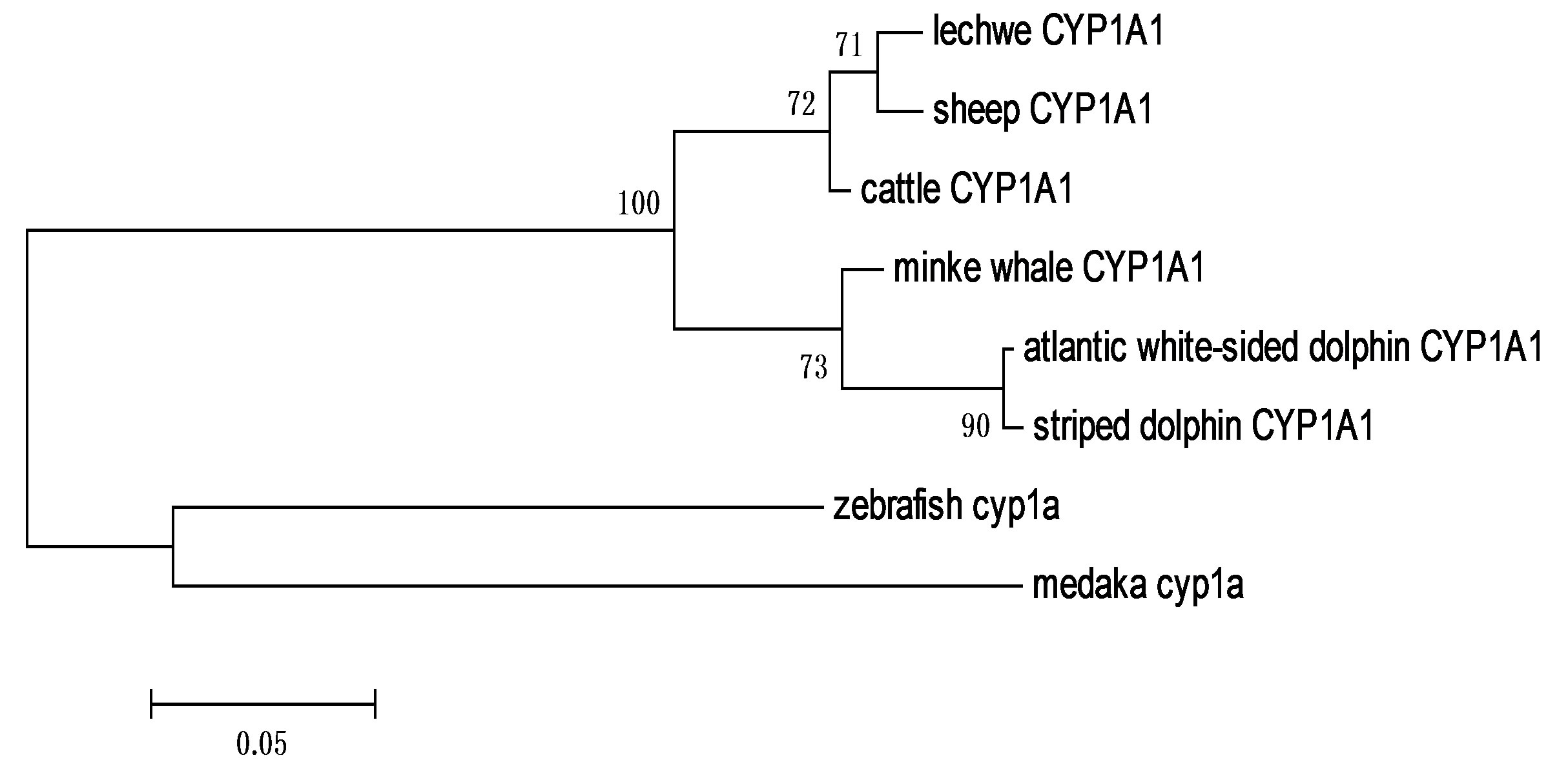
Figure 3. Phylogenetic analysis of the mammalian CYP1A1 based on the multiple alignment of the nucleotide sequences retrieved from the GenBank database. The phylogenetic tree was constructed by Maximum likelihood method. A number at each branch and the length of the stem indicate bootstrap value based on 500 samplings. Fish (zebrafish and medaka) CYP1As was used as an out-group species. CYP1A1 phylogenetic tree was corresponding to Cetruminantia phylogeny.
exposure to the pollutants in the Kafue river. Although numerous investigations have suggested that expression of CYP 1-4 isoenzymes in animals is altered by xenobiotics including environmental contaminants, and has the potential to be indicative of the exposure to these chemicals [30], there is lack of information on the characterization, sequencing and mRNA expression levels of CYP 450s in Kafue lechwe and its application. Due to the gaps in information about nucleotide and/or amino acid sequence of CYP1A1 and CYP3A in Kafue lechwe, the genes were sequenced and the results provided insights on the evolution of these genes. The partial nucleotide sequences of lechwe CY lAl and CYP3A were successfully characterized and their sequences determined and by studying the homology of the sequenced lechwe CYP1A1 with CYP1A1 of other animals, lechwe CYP1A1 showed higher similarities with CYP1A1 of sheep (98%) and cattle CYP1A1 (97%). The CYP3A lechwe sequence shared higher identities with CYP3A24 of sheep (97%) and CYP3A24 of cattle (95%). The results are in agreement with the phylogenetic analyses which confirmed the two novel lechwe genes to be part of the CYP1A1 and CYP3A24 genes. These results possibly suggest that the lechwe CYP1A1 and CYP3A24 genes could have evolved from the same ancestor with that of sheep and cattle and as such probably exhibit similar xenobiotic metabolism. The dN/dS ratio is used to infer the direction and magnitude of natural selection acting on protein coding genes. A ratio greater than one implies positive or Darwinian selection; less than one implies purifying (stabilizing) selection; and a ratio of one indicates neutral selection. The dN/dS ratio of lechwe CYP3A was estimated at 0.67 implying that this was a purifying (stabilizing) selection. This means that the genetic diversity decreases as the population stabilizes on a particular trait value. This is thought to be the most common mechanism of action for natural selection because most traits do not appear to change drastically over time. This was in agreement with the dN/dS ratio of lechwe MCR1 of 0.02. The identification and sequencing of CYPlAl and CYP3A in Kafue lechwe provided us with the invaluable information for the development of potential biomarkers of exposure to and effects of pollutants in these animals. The genes were found to be expressed in all the liver samples from both Lochnivar and Blue Lagoon GMAs (data not shown). Interestingly, CYP1A1 was higher in Lochnivar GMA, while CYP3A was higher in Blue Lagoon. A 2.9-fold increase in CYP 1A1 mRNA expression in lechwe from Lochnivar GMA compared to lechwe from Blue Lagoon GMA was seen
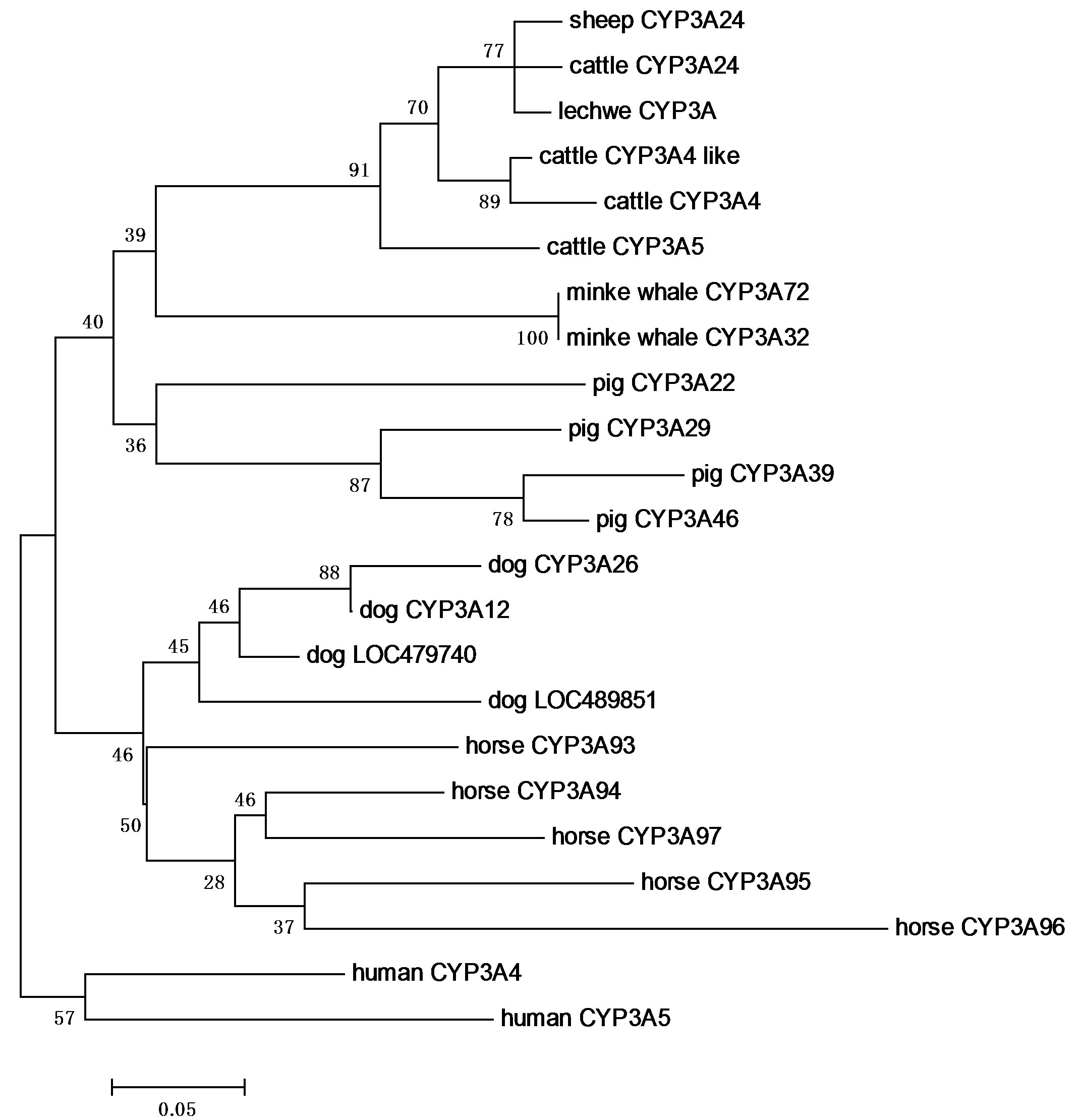

Figure 4. Phylogenetic analysis of the mammalian CYP3A based on the deduced amino acid sequences retrieved from the GenBank database by Maximum likelihood. A number at each branch and the length of the stem indicate bootstrap value based on 500 samplings. Human CYP3A4 and CYP3A5 were used as an out-group species. Novel lechwe CYP3A was clustered with sheep and cattle CYP3A24. Other CYP3As were clustered in each species.
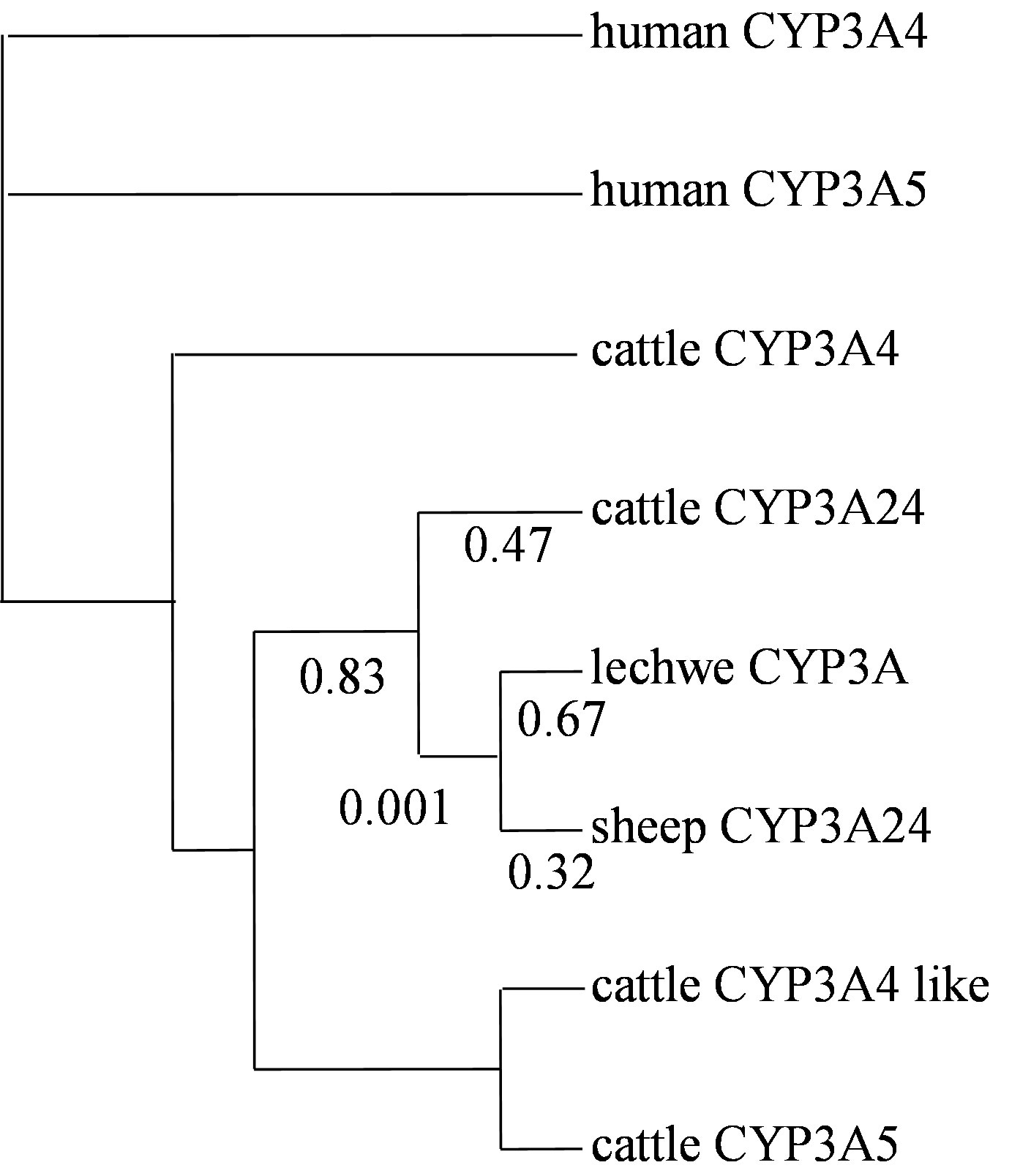
Figure 5. Phylogenetic Analysis of Maximum Likelihood (PAML) among Bovidae CYP3As. Phylogenetic Analysis of Maximum Likelihood (PAML) was used to test the differences in selective pressure among Bovidae CYP3A genes. We applied the F3x4 codon model and allowed for estimating synonymous and nonsynonymous substitution rate (dN/dS) ratio. We used 8 mammalian cDNA sequences, our novel lechwe CYP3A sequence, other Bovidae CYP3A sequences for comparing with odd toed ungulates and human CYP3A4 and CYP3A5 were used as the out-group species. The phylogenetic tree was based on genomic sequences. The CYP3A24 cluster ratios were shown.
and it could be attributed to the fact that unlike Blue Lagoon GMA, Lochnivar used to be a private ranch between 1908 and 1965 [31] a period that brackets the era when organochlorine pesticides were first synthesized and put to use for cattle dipping and other activities meant to annihilate pests in agriculture and domestic households. The Lochnivar ranch gained National Park status on the 25th of February 1972. From the time it was a ranch up to today, it has not been spared in efforts to eradicate undesirable pests and weeds by agricultural activists. Therefore it is possible that the concentration of CYP1A1 inducers in Lochnivar GMA is higher than Blue Lagoon GMA which would explain the observed higher mRNA expression in the former. A 4.2-fold increase in CYP3A mRNA expression level in Kafue lechwe from Blue Lagoon GMA compared to Kafue lechwe from Lochnivar GMA was seen. The possible reason could be that the distribution of CYP3A inducers in Blue Lagoon is higher than Lochnivar especially that Blue Lagoon GMA is comprised of cotton farmers who use a lot of DDT (CYP3A inducer) in the control of cotton bollworm.
With the different types of pesticides polluting the Kafue River and also accumulation of these pollutants in the Kafue lechwe, it is possible that the pollutants are eliciting a biological effect as seen from the mRNA expression levels determined. Therefore, CYP1A1 and
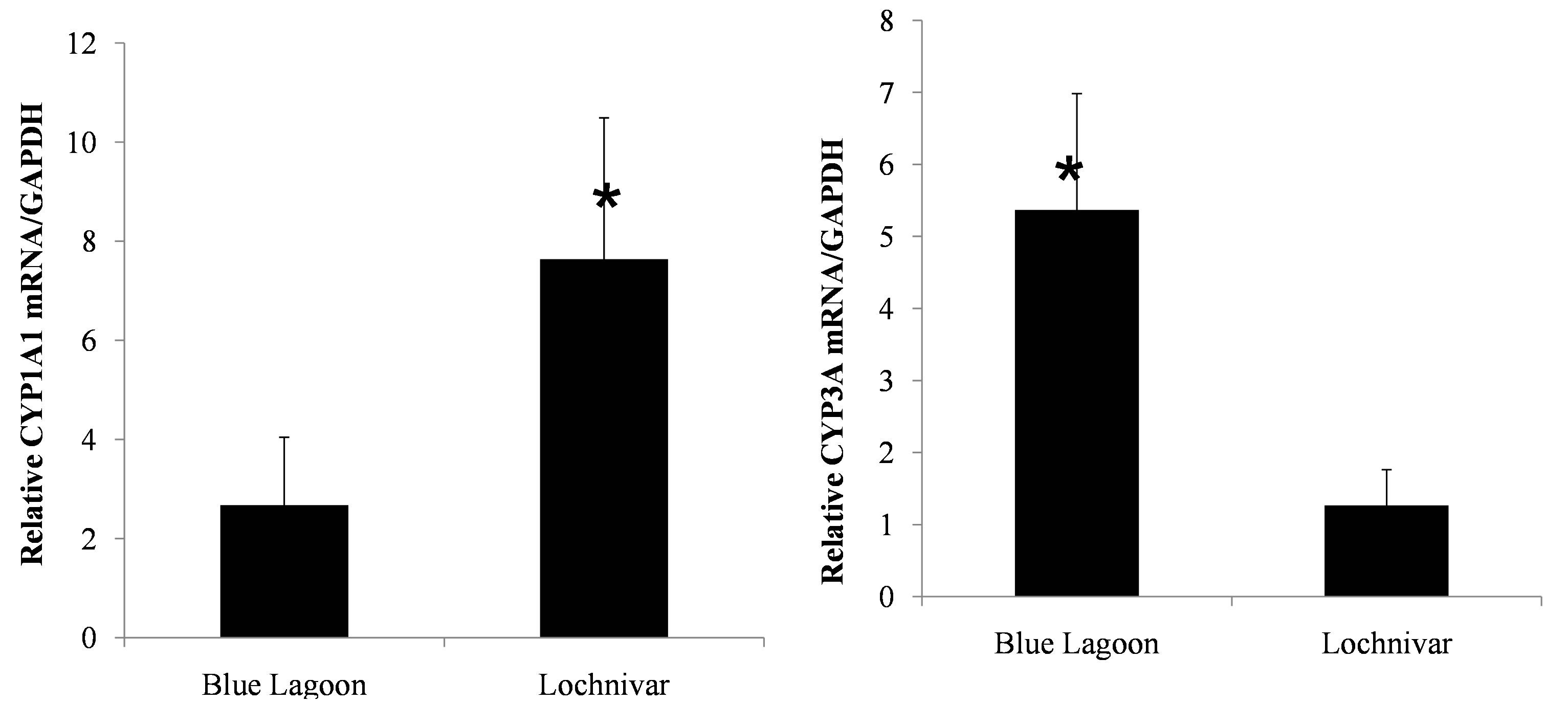
Figure 6. Regional difference of CYP1A1 and CYP3A mRNA expressions in Lechwe. *indicates significant difference (student’s t test, p < 0.05). ΔCt indicates the difference of mRNA expression levels between CYP1A1/CYP3A and GAPDH. Since there was no sex difference of CYP1A1 and CYP3A mRNA expression levels, we treated both sex as same groups. Lechwe from Lochnivar GMA showed significantly higher expression levels of CYP1A1 mRNA compared to those from Blue Lagoon GMA and lechwe from Blue Lagoon GMA showed significantly higher expression levels of CYP3A mRNA compared to those from Lochnivar GMA.
CYP3A mRNA expression levels have the potential to be used as biomarkers of pollution in these animals.
Acknowledgements
We are grateful to the members of staff at University of Zambia and Hokkaido University who participated in fieldwork and laboratory work. We thank Zambia Wildlife Authority for their cooperation and assistance. This study was supported by Ministry of Science, Technology and Vocational Training, Government of the Republic of Zambia, University of Zambia and also in part by a Grant-in-Aid for Scientific Research from the Ministry of Education, Culture, Sports, Science, and Technology of Japan awarded to M. Ishizuka (No. 24248056 and No. 24405004) and Y. Ikenaka (No. 23710038) as well as a Research Fellowship from the Japan Society for the Promotion of Science grant-in-aid awarded to S. Nakayama (No. 2403000402), and the foundation of JSPS Core to Core Program (AA Science Platforms). We also acknowledge the financial support by The Mitsui & Co., Ltd. Environment Fund.
NOTES