1. Introduction
Acute Myeloid Leukemia (AML) is a heterogeneous group of genetically diverse hematopoietic malignancies, arising from blood cell progenitors developing in the myeloid pathway or from primitive stem cells with multilineage potential. AML is understood as a complex disease resulting from multiple genetics and epigenetics alterations within hematopoietic stem cell and/or progenitor cells that have their self-renewal, proliferation, differentiation and apoptotic pathways altered. Genetic aberrations found in AML are characterized by point mutations, gene rearrangements, deletions, amplifications and a diverse array of epigenetic changes. As a general rule, gene rearrangements generated by chromosome translocations (a frequent phenomenon observed in AML) transform cells by promoting translational repression and maturation arrest (e.g. core binding factor leukemias and PML/RARA leukemias), two key events for AML burden. Such evidences support that gene rearrangements are necessary but not the exclusive event in a multistep pathogenesis raising the possibility that other contributing gene mutations occur in AML. Thus, a significant percentage of AML patients (20% - 40%) have normal karyotype and within this group and amongst other cytogenetic aberrations several somatically acquired gene mutations (e.g. key transcription factors) have been identified (e.g. NPM1, CEBPA, FLT3-ITD, TET2, WT1). The former two genes have been incorporated in the new WHO classification [1]. Till current knowledge it seems that the development of AML probably arises through a combination of genetic susceptibility and environmental factors, most of which are not yet fully understood.
Very little is known regarding to genetic susceptibility of AML. From experimental data DNA damage in the haematopoietic precursor cell is the essential prerequisite for the development of AML [2]. DNA damage can be induced directly by chemical mutagens or partially metabolized carcinogens that bind covalently to nucleic acids and proteins to form adducts. Unless priorly repaired to DNA replication, DNA adducts may lead to nucleotide substitutions, deletions and chromosome rearrangements. Chemical compounds are metabolized by a two-tiered phase detoxifying system. CYP1A1 is a member of the cytochrome P-450 phase-1 superfamily enzyme and it is a key enzyme involved in both metabolically activating and detoxifying numerous polycyclic aromatic hydrocarbons (PAH), benzo [α] pyrene (byproducts of environmental pollutants found in fuel burning, lubricant oils, fossil fuel combustion, coal, vehicle exhaust, tar and cigarette smoke) by an aryl hydrocarbon hydroxylase receptor ubiquitously expressed. PAH oxidation generates epoxide which is a very reactive compound that binds covalently DNA and unless repaired makes DNA adducts. The formation of DNA-adducts by electrophilic metabolites is generally regarded as one of the earliest steps in PAH carcinogenesis [3]. At present data, 11 CYP1A1 alleles have been reported; however, several are very rare and of unknown functional significance (a complete description can be found at http:// www.cypalleles.ki.se/cyp1a1.htm). The four most frequent polymorphisms of the CYP1A1 gene are: CYP1A1*2A a T > C at nt 6235 (T6235C) of the gene located in the 3’ untranslated region of exon 7, CYP1A1*2C, at nt 4889 in A → G transition resulting in a aminoacid change of isoleucine to valine within the heme-binding domain of exon 7; CYP1A1*3, nt 5639 at T → C transition in the 3’-non coding region, and CYP1A1*4 nt 4887 at C → A transition, resulting in aminoacid substitution of threonine to asparagine [4,5]. Phase-2 metabolism genes conjugate very water-soluble moieties like glutathione (through glutathione S-transferase system i.e. GST) to lipophilic compounds so the kidney can easily excrete the compounds. Four major classes of GSTs have been described (α, µ, π, τ) and two polymorphisms in the genes GSTM1 and GSTT1, del{GSTM1} and del{GSTT1} result in complete deletion of the gene and consequent loss of enzymatic activity. GST polymorphisms are widely spread in the population, with a large proportion of individuals presenting with homozygous deletion of the genes. There are only a few reports on AML susceptibility and it has been postulated that CYP and GST polymorphisms resulting in decreased enzymatic activity have been associated with an increased risk of cancer, including leukemia [6-8]. Also, xenobiotic metabolizing enzymes polymorphisms may influence the prognosis of AML and Hodgkin’s lymphoma, as previously shown [6]. Recently, we found out that AML patients with CYP1A1*2C polymorphisms had a better outcome when compared to wild-types alleles [9].
The genetic component of AML etiology is likely to be polygenic, and the identification of further candidate genes is essential for future studies and assessment of AML risk-related genes [10-12]. From previous studies, the results obtained are controversial and require further investigation to confirm or clarify the data obtained. We sought to determine the frequencies of carcinogen metabolism gene polymorphisms (CYP1A1, del{GSTM1} and del{GSTT1}) in a case control-study (control group and AML patients in Sao Paulo, Brazil) and predict AML risk based on polymorphism analysis.
2. Material and Methods
We studied 232 subjects, including 58 consecutively AML patients (29 males and 29 females, median age: 62 years, range 18 - 91 years) and 174 age and sex-matched control group (CG, 90 males and 84 females, median age: 50 years range 19 - 74 years). All patient samples were obtained at the time of diagnosis with informed consent at the Hematology Department at UNIFESP-EPM, Hospital São Paulo, Brazil from May 2001 to January 2004 and were stored in −80˚C until used. There were 51 de novo AML and 7 secondary AML. AML was classified according to previously standardized WHO criteria, [1, 12,13]. Thus, AML patients were categorized into three cytogenetic groups [1,12,13]. The study was approved by the institution Ethics Comittee. Samples were collected after informed consent provided, which was in accordance to the declaration of Helsinki. The control group (CG) individuals had no prior medical history of cancer and were not related to any of the patients. To verify if any of those genetic polymorphisms could pose a risk for AML there had been calculated three sex and age mathced-controls for each AML patient, according to an estimated gene polymorphism prevalence with an alpha of 0.05% (bicaudal), statistical power of 80% (beta = 0.20) with an estimated odds ratio (OR) equal or less than 2, taking into account that association with genetic polymorphism and AML would be weak. Hardy-Weinberg equilibrium was also calculated to predict frequencies of genotypes in studied population. All patients and controls were Brazilian born and lived in the city of São Paulo for at least 20 years.
Whole blood genomic DNA (from bone marrow blasts cells from AML patients and from peripheral blood lymphocytes from CG) was purified using GFX Genomic Blood purification Kit (Amersham Bioscience) following Manufacturer’s instructions. CYP1A1 polymorphisms (CYP1A1*2A, CYP1A1*2C, CYP1A1*3 and CYP1A1*4) were assessed by a PCR-RFLP assay (restriction fragment length polymorphism assays) with specific restriction enzyme digestion [4]. Polymorphisms were categorized as wild-type, heterozygous or mutant. del{GSTM1} and del{GSTT1} were studied using a PCR technique using a well standardized technique [14,15]. Beta-globin gene was amplified as an internal control. All polymerphisms categorized heterozygous or mutant were tested twice in different batches to allow prompt enzyme digestion. To verify association between case and control groups, and demographics and genetic variables Chi-square frequency test was adopted with statistical significance of 5%. Logistic regression model analysis was used to obtain relative risks or OR in order to evaluate independent prognostic factors with confidence interval of 95%. For statistical analysis STATA software, version 7.0 was used (StataCorp2001 Stata Statistical Software: Release 7.0. College Station, TX: Stata Corporation).
3. Results
Table 1 shows demographics and genotype frequencies. Genotype distributions were in Hardy-Weinberg equilibrium. Regarding to phase I detoxification enzyme polymorphisms a significant higher prevalence of the heterozygous CYP1A1*3 and CYP1A1*4 was found in AML compared to CG (65% vs 47.6%, p < 0.001 for CYP1A1*3 and 74.1% vs 66.7% (CG) p < 0.001 for CYP1A1*4). This translated into a 2.36-fold odds ratio risk for AML in carriers of heterozygous CYP1A1*3 (95% CI 1.2 - 4.5) and a 2.38-fold odds ratio risk for AML in carriers of mutated CYP1A1*4 (95% CI 0.8 - 6.8) (Table 2). On the other hand, the frequencies of the heterozygous and mutated CYP1A1*2A and CYP1A1*2C genotype were significantly lower in AML patients, than in controls (24.1% vs 61.1%, respectively, p < 0.001 for CYP1A1*2A and 22.4% vs 59.8% of controls p < 0.001, for CYP1A1*2C), reducing the AML risk to OR 0.13 for mutated CYP1A1*2A and OR 0.23 for CYP1A1*2C (Table 2). When analysing polymorphisms in phase-2 there were no differences found in del{GSTM1} frequencies between AML and CG (p = 0.999) neither del{GSTT1} frequencies (19% and 15.5%, AML respectively, and CG p = 0.539). Herein, there was no increased risk for AML in null genotypes of phase-2 genes (del{GSTM1} OR 1.0 and del{GSTT1} OR 1.27).
The sex and age-adjusted OR estimated a 2.63-fold AML risk for heterozygous CYP1A1*3 (95% CI 1.4 - 5.1) and 2.66 for heterozygous CYP1A1*4 (95% CI 0.9 - 7.8). However, heterozygous CYP1A1*2A and CYP1A1*2C were correlated to a low risk for AML (OR 0.20 and 0.19, respectively). In addition, we confirmed our data using a logistic regression model. In the multivariate analysis heterozygous CYP1A1*3 was a risk factor for AML with an OR for 3.99 (95% CI 1.9 - 8.6) whereas heterozygous CYP1A1*2A and CYP1A1*2C conferred a protective risk for AML (OR 0.54, 95% CI 0.1 - 0.6 and OR 0.23 0.1 - 0.5 respectively, for CYP1A1*2A and CYP1A1*2C) (Table 3 and Figure 1).
Karyotype was successful in 42 out of 58 AML patients. According to the cytogenetic profile 12 patients AML were categorized within the favorable group, 16 into intermediate and 14 into unfavorable (Table 1). Sixteen patients could not be cytogentically stratified due to lack of metaphases. None of the AML patients presented cytogenetics with structural deletions within the genomic location of those genetic polymorphisms studied (del1p location of GSTM1, del15q location of CYP1A1 and de l22q location of GSTT1). We did not find significant differences in polymorphism frequencies between de novo versus therapy-related AML and between different cytogenetic risk groups.
4. Discussion
We conducted a case-control study to compare frequencies of two-tiered phase detoxifying polymorphisms namely CYP1A1, del{GSTM1} and del{GSTT1}. We found a dramatic difference in heterozygous frequencies of CYP1A1*3 and CYP1A1*4 in AML patients compared to CG, whereas the frequencies of CYP1A1*2A and CYP1A1*2C were higher in CG than AML patients.
Similar findings were also described by D’Aló et al. studying Italian AML patients, in which CYP1A1*4 polymorphism presented a higher incidence in AML than in controls [16]. Thus, our results showed a higher frequency of CYP1A1*4 compared to Italians patients (18.1% vs 60.3%, respectively, p = 0.006 and p < 0.001). These differences may reflect a distinct ethnic background that may account for the occurrence of those genotypes in Italy and Brazil. The functional significance of the CYP1A1*4 polymorphism is still unknown though it might interfere with the activation of polycyclic aromatic hydrocarbons (PAH) (environmental pollutant) and estradiol metabolism.
There are no other studies reporting CYP1A1*3 frequency in AML patients [3-7,14-18]. Thus, by the multivariate analysis our data showed that CYP1A1*3 increases the risk of AML 4-fold. Till current knowledge, the association of AML risk and CYP1A1*3 genotype neither the functional significance of CYP1A1*3 was not known. The CYP1A1*3 polymorphism was reported in African-descendants and more recently in Mestizos Mexicans [4]. Moreover, the frequency of CYP1A1*3 in our control group was 47.6%. Others reports studying


Table 1. Frequency of CYP1A1, GSTM1 and GSTT1 polymorphisms among AML patients and control group.
Caucasians and Asians showed a lower CYP1A1*3 frequency. In Brazil, there is a high degree of mixed ethnic-population especially derived from Africa (former slaves) and Native Indians which could explain the frequency of CYP1A1*3 in the control group. The only reported that found out an association between cancer and CYP1A1*3 was reported by Li et al., 2004 [5]. In this study the authors verified that African-American women who had CYP1A1*3 polymorphism, smoked more than 20 years had an increased risk of developing breast cancer (OR 2.5, 95% CI 0.9 - 7.1) [5].
Surprisingly, CYP1A1*2A and CYP1A1*2C genotypes were less frequent in AML, than in controls (p < 0.001OR 0.20, 95% CI 0.1 - 0.4 and p < 0.001, OR 0.19, 95% CI 0.1 - 0.4), indicating a protective role on AML risk. This is the first report on the CYP1A1*2C polymorphism risk in AML. Henceforth we previously we published that same AML patients with polymorphic CYP1A1*2C
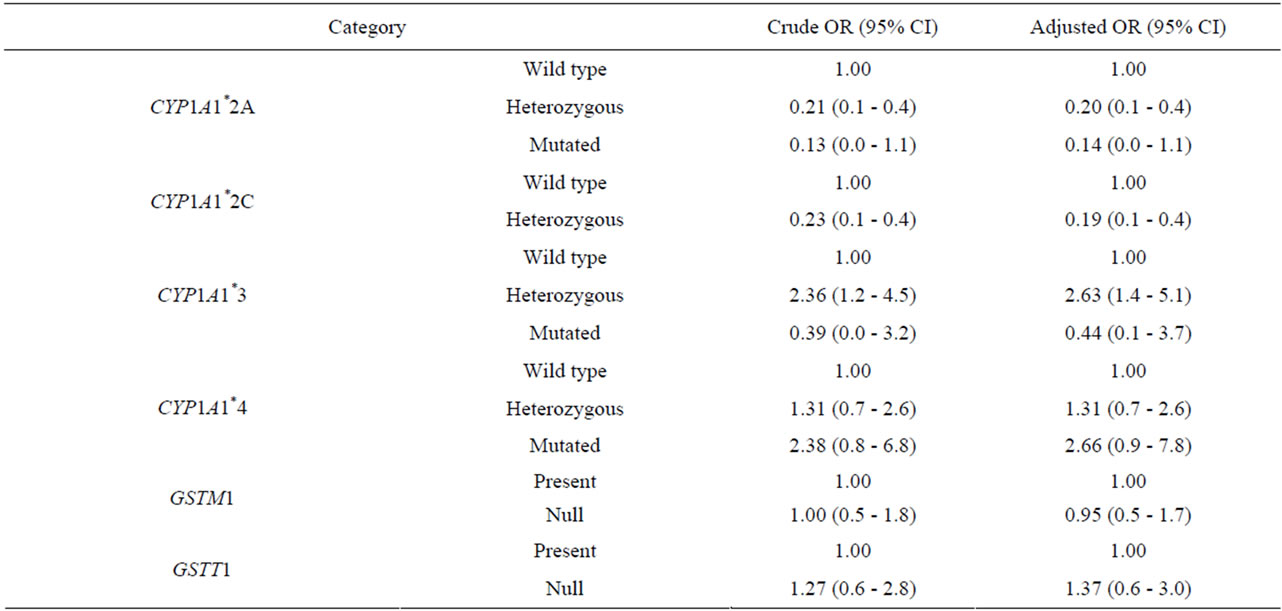
Table 2. Estimated relative risk of AML, crude and adjusted odds ratio (OR) by univariate analysis.

Table 3. Estimated risk for AML in a multivariate analysis logistic regression model.
patients had a longer overall survival compared to wild-type patients (29.2 vs 11.3 months, respectively p = 0.0261) [9]. Till current data there is no explanation for such data and further studies are required. Plus the frequency of those genotypes also vary according to different ethnic background and this may have some influence in AML risk too. Interestingly, in other medical settings such as solid tumors there is an increased risk for solid tumors in patients with CYP1A1*2A and CYP1A1*2C genotypes. Such evidence was reported in non-small cell lung cancer and head and neck cancer in Australian and Indian patients, respectively [10-12]. Experimental data from in vitro studies show that enzyme activity of CYP1A1*2C, is normal, but no further data support if this polymorphism in vivo can harbor a protective role to prevent DNA injury.
Regarding to phase-2 polymorphisms there were no differences in genotype frequencies of del{GSTM1} and del{GSTT1} in AML compared to the CG. Our findings are similar to previous reports studying Italians, British and American AML patients [6,16,17]. Those genotypes frequencies are wide spread and to establish a link with AML the cohort should be very large to have statistical significance. In another Brazilian report the frequency of del{GSTM1} was higher and harbored an increased risk of AML (OR 2.3) [17]. The patients studied might have another genotype ancestry different from our data and the number of subjects was relatively small (n = 38). We also confirmed the pattern of cytogenetic abnormalities distribution was similar to larger series of patients and previously published in Brazil and elsewhere [8,9,14,15, 17-19].
Different gene polymorphisms have proven to play a role in the susceptibility to neoplasms and it is unlikely that solely a single genetic defect might be responsible for the development of AML. According to our data carcinogen metabolism polymorphisms (CYP1A1*3 and CYP1A1*4) were correlated to a 4-fold AML risk. In which level those carcinogen metabolism gene polymerphisms might contribute to a lower detoxification carcinogen rate, bioactivation of highly intermediate toxic species, DNA adduct formation and gene mutation due to

Figure 1. Odds ratio for AML risk with regards to CYPA1*3 and CYP1A1*4 in AML patients. (a) The CYP1A1*3 polymorphism in patients with AML and the control group (CG). Blank area, wild type genotypes; gray area, heterozygous genotypes; black area, mutated genotypes. ORhet shows the odds ratio for heterozygous genotypes in a multivariate analysis; CI: confidence interval; (b) The CYP1A1*4 polymorphism in patients with AML and the control group (CG). Blank area, wild type genotypes; gray area, heterozygous genotypes; black area, negative. ORmut shows the odds ratio for mutated genotypes in a univariate analysis.
an increased level of pollutant exposure, it is still a matter of debate. Yet, it is possible that different subtypes of AML (e.g. therapy-related AML, recurrent cytogenetic abnormalities, etc.) may reflect different influences on how carcinogen metabolism polymorphisms may account for an increased DNA damage and chance to develop AML. We did not find any specific correlation within karyotypes and gene polymorphisms but we may underestimate this association due to the number of patients analyzed. Curiously, regarding to our cytogenetic data distribution is similar to larger series published elsewhere [20-22]. Others found an association with therapy-related AML and genetic polymorphisms [7]. To date, patients harboring genetic polymorphisms have an impaired ability to metabolize chemical compounds and are a higher risk suffer from occupational overexposure due to chemical agents, acquire gene mutations and develop neoplasms. It is known that benzene exposure (from industrial solvent, oil and coal emissions plants, paint Industries, automotive gasoline fumes) is a known risk factor for AML, but most of exposed people do not develop neoplasms. But how such occupational exposure may influence leukemia risk to a susceptible host (genetic variants in genes that detoxify carcinogens), it is still a matter to assess in an epidemiological study.
To the best of our knowledge this is one of the few studies assessing the risk of AML in individuals with xenobiotic metabolism polymorphisms. This is the first study to show that CYP1A1*3 heterozygous genotype increase the risk of AML. Our data support that inherited absence of this carcinogen detoxification pathway may be an important determinant of AML. Biological effects of this genotype in AML are still unknown and require further investigation. Other point to consider is the evaluation of these findings in larger series related to outcome and influence on chemoresistance.
5. Authorship
Contribution: L.A.F.P., I.D.C.C.S. and N.C.S. performed molecular analyses; M.Y. perfomed immunophenotypic analysis, M.L.L.C. performed cytogenetic analysis; and L.A.F.P., and M.L.L.F.C. wrote the paper.
6. Acknowledgements
This work was supported by the grant providedby CNPq (Conselho Nacional de Desenvolvimento Científico e Tecnológico). The fellowship grant supported by CNPq was Luís Arthur Flores Pelloso, process number 140232/ 2001-0, period 03/01/2001 to 02/28/2005.
NOTES