1. INTRODUCTION
Ginseng (Panax ginseng C. A. Mey.) distributed mainly in Changbai Mountain, Northeast of China, and which has been used as a traditional Chinese herbal medicine since ancient times [1]. In natural environment, ginseng was growing wildly, and has almost no disease happened. In the last century, with the fundament of ginsenosides have significant effect against cancers of human, requirement on it increased rapidly. And which accelerated the annihilation of wild ginseng. Now, wild ginseng was substituted by semi-wild and cultivated ginseng in the market. However, replantation of ginseng usually failed for abnormal grow, severe diseases etc., and in general, more than 30 to 40 years rotation was needed to replant successfully [2]. The following factors were suggested contribute to this problem, which including soil physicochemical characteristics deterioration, nutrient absence, severe diseases, soil microbial community unbalance and autotoxicity [3]. Many trials on fertilized application and soil sterilization have been done, but the final results was not satisfied [3,4].
Soil microorganism is one of the most important parts of soil ecosystem, and which is widely recognized as an integrative component of soil quality for their crucial involvement in many ecosystem processes. Soil microorganisms are of fundamental importance in energy and matter cycle, soil structure, soil micro-ecology balance, etc. Soil microbial communities can indirectly reflect the health status of soil, and study soil microbial communities have important significance to illustrate the mechanism of replantation obstacle. The release of root exudates, which influenced by plant species, plant age and environment factors, creates a unique and attractive environment for microbial growth and activity [5,6]. Unfortunately, the importance of rhizosphere microbial diversity is almost unknown. More recently, there is an increasing interest in the analysis of soil microbial community structure [7,8]. Among different methods, Random Amplified Polymorphic DNA (RAPD), has been widely used in species classification and phylogenetic analysis [9], species identification [10], and genetic analysis of populations [11]. As the characteristic of quick, simple and inexpensive, RAPD has become one of the most popular DNA-based methods for soil microbial community analysis [12,13].
In recent years, autoxic trail of ginseng was carried out in our laboratory, and we found methanol extract of rhizosphere soil and root exudates of P. ginseng have significant autotoxic activity [14,15]. Also, we found that, part of chemicals in the extract have allelopathy on pathogens happened on P. ginseng. So, we want to know whether microbial community in rhizosphere and nonrhizosphere soil was influenced by it. In this paper, RAPD was applied to study microbial community diversity at DNA level. The aim of the present study was to investigate the diversity of soil microbial community at rhizosphere and non-rhizosphere region of P. ginseng. Through investigate the soil microbial communities by RAPD, we expected to better understand the importance of soil micro-ecology balance and the developing mechanisms of ginseng replantation problem, and select feasible strategies to improve soil function and quality for further cultivation of P. ginseng.
2. MATERIALS AND METHODS
2.1. Soil Sample Collection
In total, thirty 1 to 6 years P. ginseng Rhizosphere and non-rhizosphere soils were sampled in field by poly-sites sampling method from Huangni village (North latitude: 42˚31'54.2" East longitude: 127˚15'45.8") and Dafang village (North latitude: 42˚26'03.2" East longitude: 127˚20'00.1") in Jilin Province of China (Figure 1, Table 1). For each rhizosphere soil sample, 5 to 8 P. ginseng roots growing at the same field were pulled up, and soil tightly adhered on the surface of root was collected. Non-rhizosphere soil samples were collected from the same depth as rhizosphere soil. All soils were put in polyethylene bags and stored at –20˚C until DNA extraction.
2.2. Extraction and Purification of Genomic DNA
Before DNA extraction, each soil sample were mixed sufficiently, and sieved (two millimeter diameter) to eliminate fibres, impurity, etc. Total microbial genomic DNA were extracted and purified by E.Z.N.A.TM soil DNA kit (Omega Bio-tek Inc.) according to the protocol. Extracted DNA were tested by 1% agarose gel electrophoresis, and stored at –20˚C. DNA templates diluted to 10 ng/µl were used for PCR amplification.
2.3. Random Amplified Polymorphic DNA Analysis
After primary screen, twenty four RAPD primers (supplied by Shanghai Sango Co. Ltd.) could amplify clearly polymorphic bands were selected from two hundred primers, and then were used to amplify soil microbial DNA in the experiment (Table 2).

Figure 1. Dendrogram of rhizosphere and non-rhizosphere soils of P. ginseng from Huangni village.


Table 1. Soil samples collected for genetic diversity analysis of microbial communities.
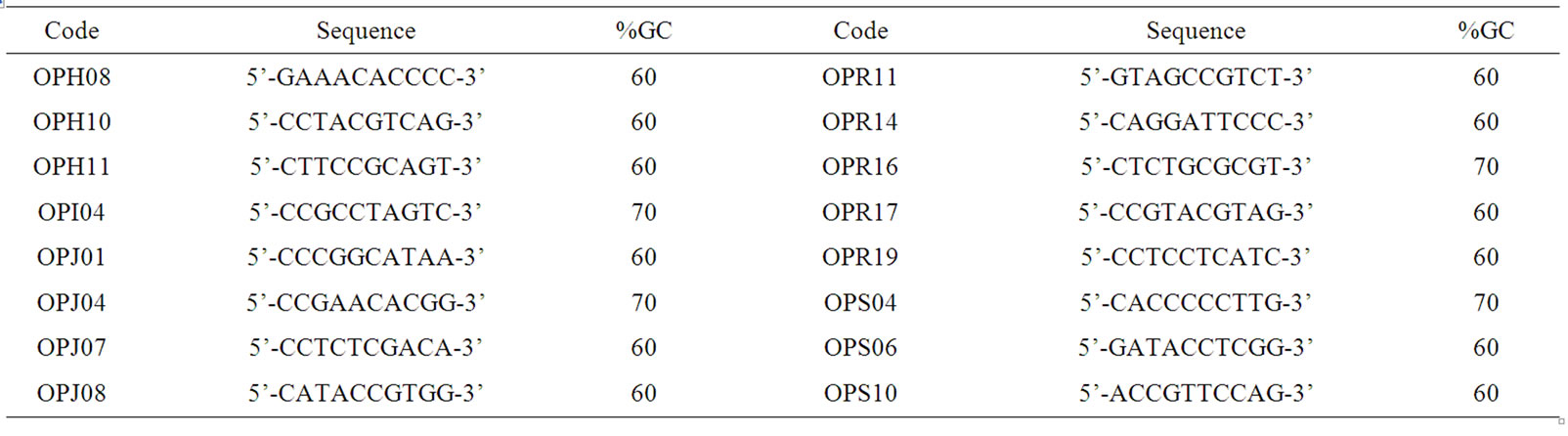
Table 2. RAPD primers used for genetic diversity analysis.
The optimized PCR volume was 25 μl, including 1 μl DNA template (10 ng/μl), 2 μl primer (10 μmol/L, Sangon Biotechnology Company Ltd., Shanghai), 1.5 μl dNTP (2.5 μmol/L, HT Biotech Ltd., Beijing), 2.5 μl 10 × PCR Buffer [100 mmol/L Tris-HCl (pH 8), 500 mmol/L KCl, 15 mmol/L MgCl2], and 0.8 μl Taq polymerase (2.5 U/μl, HT Biotech Ltd., Beijing).
Amplification was performed in a Tgradient 96 thermocycler (Biometra) with the following procedure: an initial denaturation at 94˚C for 40 s, followed by 45 cycles of denaturation at 94˚C for 20 s, annealing at 35˚C for 1 min, extension at 72˚C for 1 min, and a final extension at 72˚C for 5 min. PCR products were mixed up with 2 μl formamide buffer (98% formamide, 10 mmol/L EDTA, 0.5% bromorphenol blue, 0.5% xylene cyanol, pH 8.3), and then 6 μl of the mixture for each sample was separated on 1.5% agarose gel containing 0.5 μg/ml ethidium bromide, and photographed on gel imaging system (BioRad Cor. Ltd.).
2.4. Data Analysis
The photographic plates were scanned and analyzed using an computer-assisted image analysis system. The presence and absence of amplified bands were scored, and microbial community diversity of soil samples was measured by the percentage of polymorphic bands. Furthermore, POPGENE Version 1.32 program was employed to calculate Shannon’s information index (I) and Nei’s gene diversity (H). Similarity coefficients were calculated by NTSYSpc Version 2.10 software [16], unweighted pair group method [17] and the SHAN (sequential, hierarchical agglomerative and nested clustering) routine was used to assess the association among soils in clustering dendrogram.
3. RESULTS
3.1. Genetic Diversity of Soils Sampled from Huangni
For soils sampled from Huangni village, in total, 359 reliable RAPD fragments were generated by 24 primers. In average, 14.96 bands were amplified by each RAPD primer. Among them, 324 (90.3%) bands were polymorphic, and in average, 13.5 polymorphic bands were amplified by each primer. Statistical results showed that, bands amplified by each primer were varied from 7 to 24, and polymorphic ratio for each primer was varied from 64.7% to 100% (Table 3). Nei’s gene diversity index (H) showed that, soil sample 2 + 3N-H-G has the highest gene diversity (0.3836), and 3 + 1R-H-G the lowest (0.2373). Soil sample 2 + 3N-H-G has the highest (0.5718) ShannonWeaver index (I), and 3 + 1R-H-G the lowest (0.4005) (Table 4). And expect for soil samples (2R-H-Q and 2NH-Q) that growing 2-years P. ginseng, the Nei’s diversity index (H) and the Shannon index (I) of rhizosphere soil were lower than non-rhizosphere soil for other soil samples.
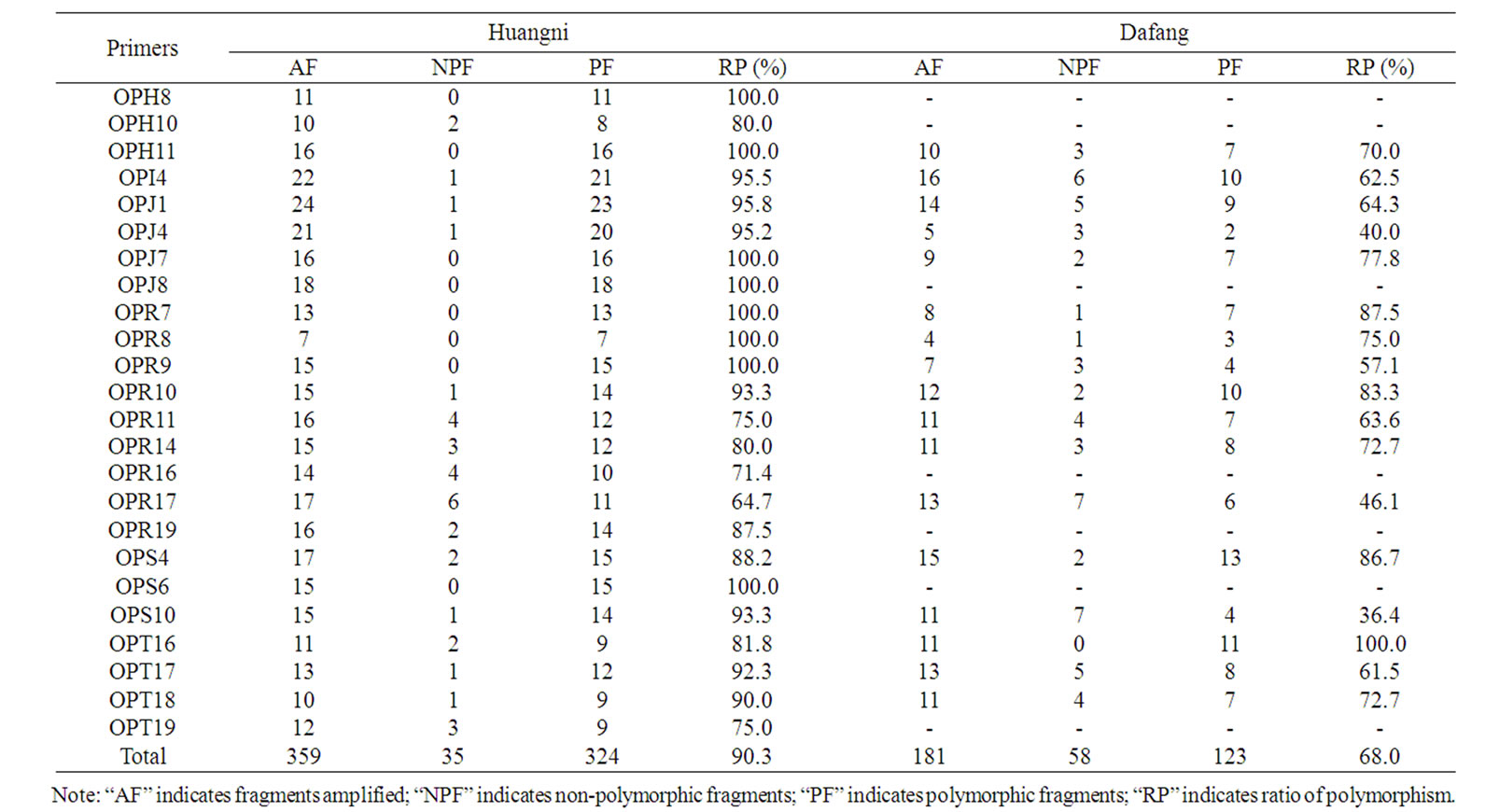
Table 3. Amplifying results of RAPD primers on microbial DNA extracted from soils sampled from Huangni and Dafang.
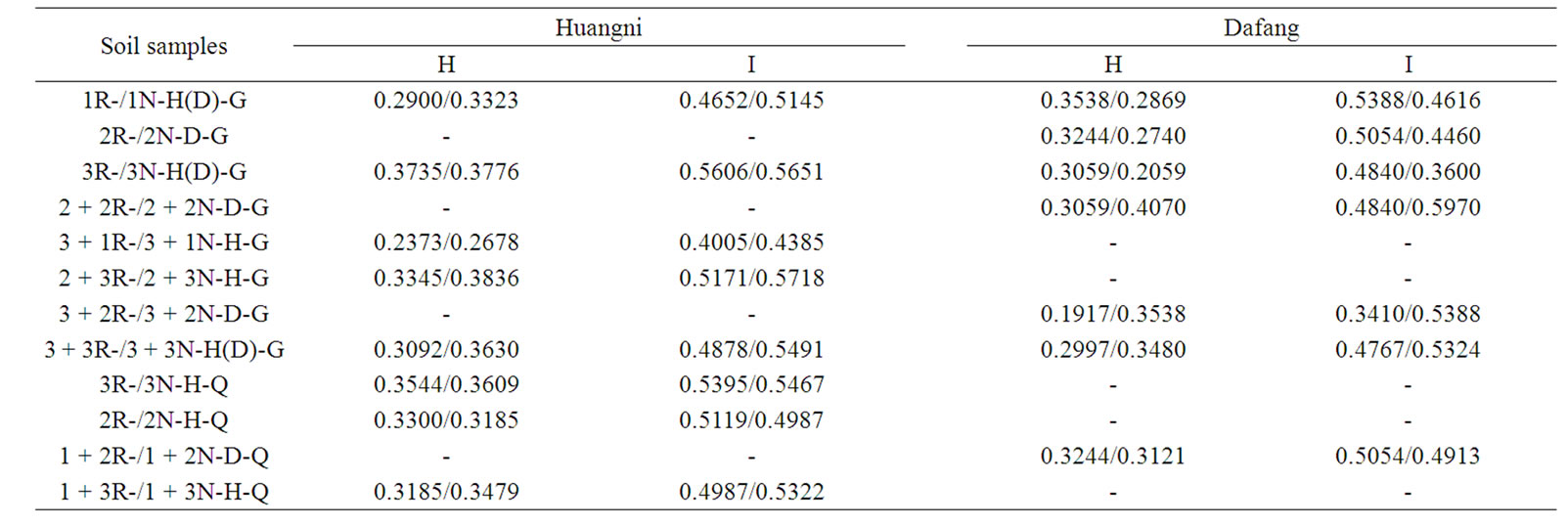
Table 4. Nei’s gene diversity and Shannon’s information index of microbial community at DNA level in soils sampled from Huangni and Dafang.
Through “0” and “1” matrix analysis by NTSYS, dendrogram were drew base on coefficients of polymorphic DNA bands (Figure 1). Results showed that, coefficient of DNA bands similarity between 3 + 1R-H-G and 2 + 3N-H-G was the lowest (0.5278), and the highest (0.8457) between 3 + 1R-H-G and 3 + 1N-H-G. Clustering results showed that, no matter P. gensing or P. quinquefolius, at the first 1 to 3 years, microbial communities between rhizosphere and non-rhizosphere soils have no significant change. For example, 1R-H-G to 1N-H-G, 2R-H-Q to 2N-H-Q, 3R-H-G to 3N-H-G and 3R-H-Q to 3N-H-Q. Though 3 + 1R-H-G and 3 + 1N-H-G were 4- years soils, which were transplanted to the new growing environment no more than one year, microbial communities showing no marked difference between rhizosphere and non-rhizosphere soils. Clustering results also showed that, no matter 4 to 6 years P. gensing or P. quinquefolius, for example, soil samples 2 + 3R-H-G to 2 + 3N-H-G, 3 + 3R-H-G to 3 + 3N-H-G, 1 + 3R-H-Q to 1 + 3N-H-Q has transplanted from one place to another growing area for a long time (about three years); In addition, the influence of root exudates on rhizosphere microbial community was increasing with the grows of P. ginseng roots, and microbial communities showed distinct change between rhizosphere and non-rhizosphere soils.
3.2. Genetic Diversity of Soils Sampled from Dafang
For soils sampled from Dafang village, in total, 17 primers generated 181 reliable RAPD fragments, in average, 10.65 bands were amplified by each RAPD primer. Among them, 123 (68.0%) were polymorphic bands, 7.24 polymorphic bands were amplified by each primer in average. The number of bands amplified by each primer varied from 5 to 16. The polymorphic ratio in each primer varied from 36.4% to 100% (Table 3). Results showed that, Nei’s gene diversity index (H) of soil sample 2 + 2N-D-G was the highest (0.4070), and 3 + 2R-D-G the lowest (0.1917). Shannon-Weaver index of soil sample 2 + 2N-D-G was the highest (0.5970), and 3 + 2R-D-G the lowest (0.3410) (Table 4). Except for soil samples (1RN-D-G and 1N-D-G, 2R-D-G and 2N-D-G, 3R-D-G and 3N-D-G, 1 + 2R-D-G and 1 + 2N-D-G) that growing 1 to 3 ages of P. ginseng, the Nei’s diversity index (H) and the Shannon index (I) of rhizosphere soil were lower than non-rhizosphere soil for other soil samples.
Dendrogram drew based on coefficient of DNA band similarity indicated differences between soil samples at DNA level (Figure 2). Results showed that, Coefficient of DNA band similarity between 1R-D-G and 2 + 2RD-G was the lowest (0.4762), and the highest (0.8730) between 1 + 2R-D-Q and 1 + 2N-D-Q. Clustering results of P. gensing soils 1R-D-G to 1N-D-G, 2R-D-G to 2ND-G showed that, in the first 1 to 2 years, no matter P. gensing or P. quinquefolius, microbial communities in rhizosphere and non-rhizosphere soil have no significant difference, and the same for soil samples 1 + 2R-D-Q to 1 + 2N-D-Q, which were the 1-year ginseng transplanted to a new growing environment no more than two years. Clustering results of soil samples 2 + 2R-D-G to 2 + 2N-D-G, 3 + 2R-D-G to 3 + 2N-D-G, 3 + 3R-D-Q to 3 + 3N-D-Q showed that, there are significant differences on microbial communities of 4 to 6 years P. ginseng rhizosphere and non-rhizosphere soil samples.
4. DISCUSSION
In this study, two hundred RAPD primers were prepared for this experiment, but only a small part of them can amplifying clearly and highly polymorphic fragments were used to reveal the influence of mono-cropping of P. ginseng on soil microbial community diversity. Results suggested that, there is significant difference on microbial community between soils sampled from Huangni and Dafang, because the number of amplified bands presents

Figure 2. Dendrogram of rhizosphere and non-rhizosphere soils of P. ginseng from Dafang village.
marked difference at two sampling sites. The Nei’s diversity index (H) and Shannon index (I) also indicated that, except for some exceptions, the microbial community diversity of rhizosphere soil was lower than non-rhizosphere soil. Dendrogram results suggested that, difference on microbial community between rhizosphere and nonrhizposhere soil of low age (1 to 3 years) P. ginseng were not significant, which was on the contrary for higher age (4 to 6 years) P. ginseng. Plants supply resources for soil communities by providing organic matter through leaf-litter inputs, through the release of root exudates, or through other ways of deposition of organic compounds into the soil environment, and as a result, specific soil communities form under different plant species [18,19]. So we deduced that, as the primary carbon source for the sustentation of soil microbial community, root exudates have directional selection press on microbial species growing in the soil environment, and finally resulted in the degradation of microbial communities.
Usually, rotation provides a greater concentration and diversity of organic materials than the mono-cultural systems, which can improve microbial community diversity and stabilized microclimate to perform their function [20,21]. The RAPD analysis demonstrated that there is significant diversity on not only microbial communities between rhizosphere and non-rhizosphere soils of P. ginseng or P. quinquefolius, but also plants with different ages. Furthermore, Nei’s gene diversity and Shannon’s information index in soils were influenced by mono-cropping significantly. In this present, the difference on microbial community diversity is increasing with the growing years of P. ginseng at a given site. And, the influence was more notable for 4 to 6 years ginseng. Result demonstrated that, mono-cropping systems have negative effects on soil microbial communities, and with the growing of P. ginseng root, the influence tend to be more and more marked. On one hand, mono cultivation systems decrease given organic matter which is benefit to soil microorganisms, and made diversity of microbial community decrease. On the other hand, the environmental factors, such as overuse of nitrogen, phosphorus fertilizers, germicide and pesticides, exudates released from root to their adjacent soil may induce changes of soil microbial community and soil physic-chemical properties, also [13,22].
Historically, the research on biodiversity was mainly concentrated on plants and animals, despite microorganisms being the critical components in the soil ecosystems, microbial diversity was regarded as an indicator of soil quality, and study on their diversities was hampered due to the difficulties in measuring. Since no more than 10% soil microorganisms have been isolated and studied yet [23]. Furthermore, the spatial distribution and interaction may have influence on the quantitative and representative of microorganisms in soil [24]. Molecular methods are emerging as tools to better study microbial communities in soils [25]. The difference among microbial species is essentially represented in their genetic diversity at DNA sequence. Thus, soil microbial community diversity can be studied by using molecular techniques at DNA level.
RAPD markers require no genomic structure information on the materials studied, and the binding sites of which are randomly distributed on the genome DNA; furthermore, it is easy-operating, economic, effective and yields a great deal of information to discover small variations between similar genomes [10,26,27], all these made the RAPD technique an popular DNA-based method for assessing soil microbial community. In this study, RAPD technique revealed the influence of continuous P. ginseng cultivation on soil microbial community diversity. The results may suggest that the change in microbial community DNA composition was accompanied with the change in microbial functional properties.
However, the same as other DNA-based techniques, such as DGGE, ARISA, T-RFLP, etc., RAPD technique also has its limitation for determining soil microbial community. As that RAPD primers only 10 oligonucleotide bases, many factors involved in the amplification could affect the reproducibility and instability of RAPD results [28,29], and which made the repeatability of RAPD not as perfect as other PCR-based methods. In order to determine the most suitable RAPD conditions, concentration and proportion of Mg2+, dNTP, template DNA and primer, annealing temperature and extending time were evaluated. Another problem worth attention is that, as other PCR-based methods, RAPD amplification was also sensitive to the concentration of DNA templates, only those more than 1% of total DNA can be detected by this method [30]. Although the effects of PCR bias, RAPD method was still used in genetic diversity research [31,32]. However, as the enormous of microorganisms and the limited resolving ability of RAPD, it is necessary to integrate diverse approaches and perspectives to understand more precisely the changes in the diversity of microbial communities [33,34].
5. ACKNOWLEDGEMENTS
The authors thank Mr. H. M. Fang and Mr. Q. Y. Gao for assistance collecting soil samples. The work was supported by a grant from the National Natural Science Foundation of China (81072992) and Doctoral Fund of Ministry of Education of China (200800231060).
NOTES