Analysis of Some Important Forage Quality Attributes of Southeastern Wildrye (Elymus glabriflorus) Using Near-Infrared Reflectance Spectroscopy ()
1. Introduction
The southeastern wildrye was originally described as a variety of Canada wildrye (Elymus canadensis L.) [1] . At times, it was also classified as a variety of Virginia wildrye (Elymus virginicus L.), but more recently it is recognized as a distinct species (Elymus glabriflorus (Vasey ex L.H. Dewey) Scribn. & C.R. Ball) [2] . Southeastern wildrye is a native, tall (around 4 foot), cool season, clump-forming, perennial grass. Belt et al. [1] described southeastern wildrye as a species that is tolerant of a wide range of growing conditions from moist to dry soil, partly shade to full sunlight, coarse to fine textured soils, early successional to mature meadow communities, and acid to neutral soil pH. Naturally, it is found in woodland edges, open woods, temporary and permanent meadows, thickets and open grasslands, sometimes spreading into old fields and roadsides [2] . Recently, this grass has gained a considerable interest in its development as a forage crop for the Southeastern USA. Tall fescue (Schedonorus arundinaceus (Schreb.) Dumort., nom. cons.) is the only other perennial cool-season forage crop that can be incorporated into grazing and haying systems in this region of the country.
In the southern edges of the southeastern region, annual grasses such as ryegrass (Lolium multiflorum L.), oats (Avena sativa L.), and cereal rye (Secale cereal L.) are used for cool-season grazing options, whereas tall fescue does not persist. However, southeastern wildrye, due to its wide areas of adaptation, grows very well across the whole southeastern region of the US.
Rushing [3] observed southeastern wildrye as a good cool-season forage, with high crude protein (13% - 19%), low neutral detergent fiber (45% - 55%), and low acid detergent fiber (255% - 35%) contents with dry matter yields of more than 3 tons per acre on unfertilized stands. Rotational grazing is recommended in order to maintain forage quality (harvest or graze every 20 - 30 days) and quantity. High innate nutritional content makes southeastern wildrye a great cool-season option for those interested in a year-round grazing system using only native species [1] . In evaluating southeastern wildrye as a forage crop, it has become necessary to analyze this species in field trials where forage quality attributes (nutritive values) must be quantified. Forage quality attributes are essential criteria in the selection, management, and improvement of forage for livestock consumption [4] . The protein and detergent fiber (ADF and NDF) contents are the primary forage quality attributes that determine forage intake and digestibility, which are considered as the two main factors of ultimate forage quality [5] .
As far as abundance of various elements are concerned, C, H, and O constitute about 99.9% of the dry matter of most plant biomass with H being the most abundant atomic species [6] [7] . This indicates that the chemical bonds in organic compounds in plant biomass predominantly involve C, H, and O. Transmittance or reflectance of light in the NIR region (700 - 2500 nm) is known to interact with functional groups frequently found in plant biomass. These include the groups such as O-H as in water, C-H, C-O, C-O-H, N-H, and C=C bonds, which cover most of the covalent bonds of various organic compounds exist in plant biomass, except for the C-C single bonds in the carbon chains [8] . This suggests that NIR spectroscopy (NIRS) can be a viable, non-destructive, rapid technique for analyzing various quality attributes of feedstuffs including grasses. For example, NIRS has been successfully used to predict the nutritive value of forages and hays through direct scanning of the forage samples or the extrusa obtained from esophageally fistulated animals [9] - [17] . Several other reports have also demonstrated successful application of NIRS in the analysis of anti-quality factors in forage [18] - [20] . Vogel et al. [21] developed NIRS calibration models to determine over 20 different quality attributes of switchgrass including cell wall and soluble sugars. Thus, a non-destructive spectroscopic sensing technique has the potential to be an accurate method to rapidly determine the forage quality of southeastern wildrye, which is a potential new forage crop for both grazing and haying in the southeastern United States. However, there is no such work done on developing useful NIRS calibration models for routine analysis of the forage quality of southeastern wildrye, which has a great potential as forage crop both in terms yield and quality.
Like other new crops with specific niches for production, like switchgrass, a properly developed and validated NIRS calibration equation for southeastern wildrye would provide a very useful tool to rapidly compare the forage quality of different varieties grown under various management practices under variable soil and climatic conditions and harvested differently with respect to age, cutting/grazing height and others. In this study, we successfully developed and validated NIR spectroscopic calibration models for dry-matter, crude protein, acid detergent fiber, neutral detergent fiber, digestible dry-matter, dry-matter intake, and relative feed value of southeastern wildrye, which are considered as the principal attributes of forage quality.
2. Materials and Methods
2.1. The Samples and Sample Preparation
Samples used to develop and validate NIRS calibration equations in this study were obtained from three field trials conducted during 2011 and 2012. Two of three field trials were conducted at the H.H. Leveck Animal Research Farm (South Farm) on the campus of Mississippi State University. The first trial was a perennial cool-season variety trial in which 18 species were evaluated. These included seven wildrye species, two wheatgrasses, five orchardgrasses, one timothy, and three tall fescues. This study was a randomized complete block design with four replicates. Plots were established in the fall of 2010. Plots were fertilized according to soil test recommendations, however no nitrogen was applied. A total of seven harvests were made (four in 2011 and three in 2012). The second study at this site was another cool-season variety trial, however seedlings were grown in greenhouse conditions and out-planted in a randomized complete block with four replicates. In this study, the same seven wildryes and one tall fescue (“Kentucky-31”) were evaluated. As with the other study at this location, no N was applied. A total of four harvests were completed for this trial (one in 2011 and three in 2012). The third study was conducted at the R.R. Foil Plant Science Research Center (North Farm) on the campus of Mississippi State University. For this field trial, southeastern wildrye was sown in 10 foot rows in the fall of 2010. The trial consisted of five harvest interval treatments (20, 30, 40, 60, and 80 days). Fertilizer was applied at a rate of 50 l b of N using 15-5-10 (N-P2O5-K2O) prior to each spring’s harvests. For all studies, sub-samples were taken for each plot or row for each harvest. Though various cool season perennial grass species were included in these trials and samples of all species were collected, only southeastern wildrye samples were used for this work. Samples were weighed, placed in a forced-air oven and dried at 65˚C until no further change in weight could be measured. Samples were ground using a Wiley-type mill with a 1 mm screen and stored at room temperature until used for laboratory analyses of dry-matter, crude protein, ADF, and NDF contents as well as for collecting NIR spectra as described below.
2.2. Determination of Dry Matter Content
Dry matter content of the pulverized samples was determined as per the Association of Official Analytical Chemist (AOAC) method 930.15 [22] by drying approximately 2 g of sample in a forced-air drying oven at 135˚C ± 2˚C for at least 2 h with freely circulating air. Aluminum dishes with covers were dried at the aforementioned temperature and time; then, they were placed in a desiccator and allowed to cool to room temperature. Tare weights were obtained for containers with covers (W4) to the nearest 0.1 mg. Approximately 2 g of sample were added to each container. Total weight was recorded (W5) to the nearest 0.1 mg. Dishes were shaken to gently distribute sample and expose maximum area for drying. The dishes, with sample (uncovered) sitting on the covers, were placed into preheated oven at 135˚C and allowed to dry for 2 h. Samples were then placed in a desiccator, covered, sealed, and allowed to cool to room temperature. The dish was weighed with covers (W6) and recorded to the nearest 0.1 mg. Percent dry matter (DM) on wet basis was calculated using the following equation:
Calculation: 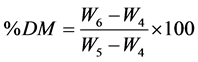
where: W4 = tare weight of dish in grams.
W5 = initial weight of sample and dish in grams.
W6 = dry weight of sample and dish in grams.
2.3. Determination of Crude Protein Content
Crude protein content of the samples was analyzed based on Total Kjeldahl Nitrogen (TKN). The laboratory procedure for TKN included a digestion step followed by a distillation-titration step. Procedures followed the Association of Official Analytical Chemist (AOAC) method 984.13.15 [23] . For each batch, a glutamic acid sample was digested and analyzed as a quality control (QC) standard (i.e., a certified reference material). Besides, each batch included one each of a blank, laboratory control spike (spiked on blank), matrix spike (spiked on a randomly chosen sample), and duplicate (of a randomly chosen sample). These QC samples were digested for TKN and subsequently analyzed for NH4-N. The acceptance criteria for these QC samples were as follows: Blank, below the method detection limit; QC standard, recovery within 80% - 120%; laboratory control spike, recovery within 80% - 120%; matrix spike, recovery within 70% - 130%; duplicate, relative percentage difference ≤15%.
Approximately 1 g of ground sample, weighed to nearest 1.0 mg (W), was placed into a 250 mL TKN digestion tube. A solution comprised of a copper catalyst of 15 g K2SO4 and 0.04 g anhydrous CuSO4, 3 g of pumice, and 20 mL of concentrated sulfuric acid was added to the flask. The tubes were swirled and placed in a metal rack. The digestion tubes along with the metal rack were then placed into the unheated digestion block (BD-20, Fisher Scientific, Inc., Pittsburgh, Penn.). As the first step, the digestion block was initially brought stepwise to a temperature of 200˚C, then to 300˚C, and finally to 420˚C. The samples were then digested for 90 minutes or until the samples were green and clear. The tubes with digested samples were removed from the block and cooled for 15 min at room temperature and diluted by addition of 100 mL deionized water. The Kjeldahl digests were distilled using an autodistillation unit (Kjeltec 8100, Foss-North America, Eden Prairie, MN). The method used a 40% NaOH solution to generate an alkaline distillation environment for producing ammonia vapor and 2% boric acid (4% for the glutamic acid digest) solution to collect the distilled ammonia. The boric acid solution also contained 0.001% bromocresol green and 0.0007% methyl red indicators to indicate the endpoint during subsequent titration with standardized hydrochloric acid. The collected ammonia in boric acid solution was titrated with 0.2 N HCl to the endpoint and volume of titrant consumed was recorded to the nearest 0.1 mL (VA). The titrant required to titrate the reagent blank (VB) was also recorded. Percentage nitrogen was calculated using the following equation:
Calculation: 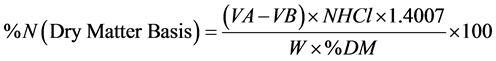
where:
VA = volume (mL) of standard HCl required for sample
VB = volume (mL) of standard HCl required for blank
NHCl = normality of standard HCl
1.4007 = milliequivalent weight of N × 100
W = sample weight in grams
Percentage crude protein (CP) was calculated using the following equation:
%CP (Dry Matter Basis) = %N (Dry Matter Basis) × 6.25.
2.4. Determination of Acid Detergent Fiber (ADF) and Neutral Detergent Fiber (NDF) Contents
The Acid Detergent Fiber (ADF) and Neutral Detergent Fiber (NDF) contents were determined from 0.5-g pulverized samples using an automatic Fiber Analyzer (ANKOM2000 Fiber Analyzer, ANKOM Technology, Macedon, NY) based on the methods described by Goering and Van Soest [24] and Van Soest et al. [25] . The solvent used for the ADF was an acid detergent solution (FAD20C, ANKOM Tech.). The solvent used for the NDF analysis was a premixed neutral detergent solution (FND20C, ANKOM Tech.), which contained 30.0 g sodium lauryl sulfate, 18.61 g ethylenediaminetetraacetic acid disodium salt dihydrate, 6.81 g sodium tetraborate decahydrate, 4.56 g an hydrous sodium phosphate dibasic, and 10.0 ml triethylene glycol dissolved in 1 L distilled water with pH within 6.9 to 7.1. Twenty grams sodium sulfite and 4.0 ml of heat stable alpha-amylase with activity = 17,400 Liquefon Units/ml (catalog #FAA) were added to 2000 ml of ND solution used to process 24 sample bags. Filter bags were suspended in the reaction vessel in a stainless steel suspension basket. The samples were processed in the reaction vessel for 70 min for the ADF procedure and 80 min for the NDF procedure. The solution was then drained and the vessel was filled with 2 L of 94˚C deionized water and the samples were agitated for 5 min. The hot water rinse was repeated 4 times. The filter bags were then removed and placed in a small beaker and covered with acetone to soak for 5 min. The bags were then squeezed and dried in an oven at 100˚C. The NDF and ADF concentrations were calculated using the following equation:
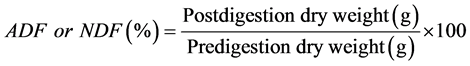
2.5. Computation of Digestible Dry Matter (DDM), Dry Matter Intake (DMI) and Relative Feed Value (RFV)
The DDM, DMI, and RFV were calculated as follows [26] :
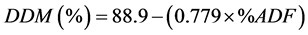

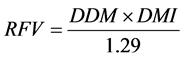
2.6. Near-Infrared Reflectance Analysis
Scanning of the samples was performed using a NIRSystem model 6500 near-infrared scanning monochromator (FOSS North America, Eden Prairie, Minnesota) in the reflectance mode. The instrument was equipped with a combination of silicon and lead sulfide detectors. Following instrument warm-up, satisfactory instrument performance was confirmed through instrument response, photometric repeatability (noise), wavelength accuracy tests, and check cell scan. For the analysis, subsamples of the homogenized pulverized samples were packed in ring cups (Part# IH-0386, FOSS North America, Eden Prairie, Minnesota) as follows. The cup was first overfilled, and the excess was removed by scraping it away. This procedure resulted in approximately 5-g samples being scanned (around 10 mm depth). The packed cup was held on a transport module and 32 successive scans were carried out covering the wavelengths from 400 to 2498 nm at 2-nm intervals to give a 1049 data points per sample. For the control, 16 scans were made over an internal standard ceramic disk before and after the samples were run. The reflectance energy readings were referenced to the corresponding readings from the internal standard. An average of 32 successive scans was used to record the spectrum of each sample. All spectral data were recorded as the logarithm of the reciprocal of reflectance (log 1/R, R = reflectance). The scanning procedure was completed in 1.5 minutes per sample and this excludes the time taken for instrument warm up and performance check.
Absorption of radiation at 400 - 2498 nm wavelength, the visible plus near-infrared region, was used to develop calibration equations related to the dry-matter (DM), crude protein (CP), acid detergent fiber (ADF), neutral detergent fiber (NDF) and digestible dry-matter (DDM) contents, and dry-matter intake (DMI), and relative feed value (RFV) of the samples. After collecting the NIR spectra of all 179 samples, mathematical processing, and statistical analysis were performed using the software package WinISI WinSCAN v1.50 (FOSS North America, Eden Prairie, Minnesota). In WinISI software, the score program was used to select samples for spectral outliers before calibration and validation. For scoring, the principal components regression (PCR) analysis of the sample spectra was used. The score algorithm ranked spectra according to Mahalanobis distance. The first Mahalanobis distance accounted for difference of a given sample spectrum from the average spectral feature of the sample set as a whole [27] , called “global-H” or “GH” distance. This gives spectral boundaries to eliminate the outlier samples with GH > 3.0. The algorithm also measured similarities between sample spectra with a second Mahalanobis distance, called “neighboring-H” or “NH” distance and eliminates samples with NH < 0.6 from their nearest neighboring samples. Elimination of samples based GH and NH distances assisted in the development of an accurate and robust prediction equation [28] . After elimination of spectral outliers, the qualified sample sets for each of dry-matter (DM), crude protein (CP), acid detergent fiber (ADF), neutral detergent fiber (NDF) and digestible dry-matter (DDM) contents, and dry-matter intake (DMI), and relative feed value (RFV) of the samples were randomly divided into two subsets: two-third for calibration and cross validation, and one-third for external (or independent) validation to test the performance of the developed equations. The external validation sample set allowed the NIRS equation to be validated for prediction accuracy, using random samples not included in the calibration sample set [29] .
Development of calibration models and validation of the developed models were performed using the global program in WinISI software as outlined in the WinISI software manual using wavelengths of the entire visible (400 - 1100 nm) and near-infrared (1100 - 2498 nm) regions at every 2 nm interval. For calibration model development, we used a linear regression method, which was based on a modification of the partial least-squares (PLS) algorithm [27] , where the X and Y residuals are standardized at each iteration, called “modified partial least square (MPLS)” regression. Pretreatments with standard normal variate and detrending (SNVD) were used for scatter correction. The SNVD was designed to remove additive baseline and multiplicative signal effects to give a spectrum with zero mean and a variance equal to one. Application of SNVD transformation to raw spectral data reduces the interference of physical characteristics such as particle size and path length of sample to the spectra [30] [31] . The optimum number of PLS loading factors (called terms) was determined by cross-validation of MPLS procedure, and it was 6 for all models. Calibration models were optimized applying second derivative treatment of the spectra and 4-nm gap (i.e., 8 nm, the spacing over which the derivative was calculated). Only the first smoothing (the number for data points in a running average) at 4 was set in the software avoiding the second smoothing option. The application of the second-derivative algorithm to the raw spectra (log 1/R) resulted in an increase in the complexity of spectra and a clear separation between peaks, which overlaps in the raw spectra [32] .
The calibration development procedure included a cross-validation of the calibration models. The cross-validation procedure of WINISI software estimated the validity of the calibration models by selecting every fifth sample in the calibration set and holding it for use as a validation during calibration. Thus during the cross-validation procedure, each calibration subgroup included 80% of the samples and the validation subgroup included the remaining 20%. The very first cross-validation set was selected using the first sample as a starting point. The second cross-validation set was selected using the second sample as a starting point. In this manner, the cross-validation procedure was performed at least five times so that every sample in the entire set was used in the validation procedure. Thus, the procedure did not compromise the robustness of the calibration models by holding back a set of samples for validation. During each cross validation step, the model outliers were rejected based on their spectral differences (H statistic) as described above. Such internal cross validation avoided overfitting of the equations by selecting the minimum number of PLS terms in each model [33] .
Once the models were developed, a further elimination process removed the compositional outliers from the calibration sample set on the basis that the difference between predicted and laboratory-measured values exceeded three times original SECV [34] [35] . Compositional outliers are believed to be the samples with poor quality laboratory-measured values that do not correlate well with the spectral features of the samples [34] - [36] . After exclusion of the compositional outliers, the final calibration models were developed, which were able to give NIR-predicted values within three standard deviations from the mean difference when compared with the associated laboratory-measured values for each sample included in the model.
Quality of the developed calibration models were evaluated based on the lower standard error of calibration (SEC) and higher coefficient of determination for calibration (R2) that accounts for the proportion of explained variation by the model. The cross validation ability of the models was evaluated based on the overall standard error of cross validation (SECV) and the associated 1-variance ratio statistics (1-VR) derived from the outcomes of all five cross validation steps. The 1-VR is indeed the coefficient of determination in cross validation. Besides, the following two ratios were utilized to evaluate the quality of the models [37] [38] :
(1) RSCD, SD ÷ SECV, the ratio of standard error of cross validation to deviation (SD, standard deviation of reference data in calibration set).
(2) RSCIQ, IQ ÷ SECV, the ratio of standard error of cross validation to inter-quartile distance (IQ, inter-quartile distance in reference data in the calibration set).
The DM, CP, ADF, NDF, DDM, DMI and RFV of sample sets, randomly separated out for external validation, were predicted by the developed models and compared with the associated laboratory-measured values. Once the samples were predicted, the compositional outlier samples were also removed from the validation set if the difference between predicted and laboratory-measured values exceeded three times original SECV [34] [35] . After exclusion of such compositional outliers, all remaining samples in the validation set gave NIR-predicted values within three standard deviations from the mean difference when compared with the associated laboratory-measured values. The standard error of prediction (SEP), bias-corrected SEP [SEPC], and the associated R2 were then used to evaluate the prediction performances of the models. Besides, the following two ratios were utilized to evaluate the success of external (independent) validation of the models [37] [38] :
(1) RPD, SD ÷ SEP, the ratio of performance (SEP) to deviation (SD of the reference data in the external validation set).
(2) RPIQ, IQ ÷ SEP, the ratio of performance (SEP) to inter-quartile distance (IQ of the reference data in the external validation set).
3. Results and Discussion
3.1. Spectroscopic Analysis
An average raw NIR reflectance spectrum of the wildrye samples are shown in Figure 1(a). The second derivative was calculated from the log (1/R) spectra at gaps of 4 data points (8 nm) and a smoothing over segments of 4 data points (2, 4, 4, 1) with scatter correction (SNVD) and the derivative form of an average spectrum is shown in Figure 1(b).
In the average raw spectrum (Figure 1(a)), the main absorption bands were observed over several wavelengths. The absorption band covering 1436 - 1464 nm was related to C-H combination bands of methylene (−CH2) group associated with aliphatic and aromatic hydrocarbons, C-H stretching first overtone of starch, N-H stretching first overtone of amide/protein, N-H stretching (symmetric) first overtone of aromatic amines, >C=O stretching third overtone of ketones and aldehydes, and O-H stretching first overtone of water, starch or polymeric alcohol. The 1720 nm band was related to C-H stretching first overtone of methylene (−CH2) group associated with aliphatic and aromatic hydrocarbons, S-H stretching first overtone of thiol group associated with thiols and the band at 1926 nm was related to O-H stretching and HOH deformation combination from molecular water or water molecules in the 3-aminopropyltriethoxysilane-ethanol-water system, O-H stretching and HOH bending combination in polysaccharides, and >C=O stretching first overtone in amides. The band covering 2100 - 2136 nm was related to O-H bending or C-O stretching of starch, C-O-O stretching third overtone of starch or cellulose, whereas the band covering 2302 - 2344 nm was related to C-H bending second overtone of proteins, C-H stretching and CH2 deformation combination from starch and other polysaccharides, C=O hydrogen bonded to the N-H of the peptide link (termed the α-helix structure) of proteins, and C-H stretching (symmetric) first overtone of methylene group associated with aliphatic hydrocarbons, and C-H bending from polysaccharides. The band at 2488 nm was related to C-H stretching and C-C combination of cellulose. The overlaid raw spectra for all 179 samples shown in Figure 2 reflect the fact that they belong to same population.
The second-derivative spectra had a trough corresponding to each peak in the original spectra, removing the overlapping peaks and baseline effects [39] . The second derivative of an average spectrum (Figure 1(b)) showed absorption bands at 1398 nm related to C-H bands of methylene (−CH2) group associated with aliphatic and aromatic
![]()
![]()
Figure 1. Raw spectrum (log 1/R; (a)) and second derivative (2, 4, 4, 1 + SNVD; (b)) of NIRS average spectrum of southeastern wildrye. “R” stands for reflectance; “SNVD” stands for “Standard Normal Variate and Detrending”; “2, 4, 4, 1” stands for second derivative, 4 data points for gap, 4 data points for first smoothing, and avoidance of using second smoothing. For further details, see Materials and Methods section.
![]()
Figure 2. Overlaid raw spectrum (log 1/R) of all 179 southeastern wildrye samples. “R” stands for reflectance.
hydrocarbons; 1454 nm related to O-H stretching first overtone of starch, C-H bending of methylene (−CH2) of hydrocarbons and O-H stretching of starch; 1480 nm related to N-H stretching first overtone of −NH2C=O; 1526 nm related to N-H stretching first overtone of −NH2R; and 1742 nm S-H stretching first overtone of thiol group associated with thiols. The bands at 1780 nm was related to C-H stretching first overtone of cellulose and H-O-H deformation and bending of cellulose, 1794 nm was related to O-H bending of water, 1962 nm was related to O-H stretching and O-H bending combination band of starch, 2099 nm was related to O-H bending or C-O stretching third overtone of starch or cellulose, 2186 nm was related to N-H bending second overtone of protein, C-H stretching or C=O bending of protein, 2280 and 2348 nm were related to C-H stretching or −CH2 deformation-bending of starch, and 2304 nm was related to C-H bending second overtone of protein.
The NIRS, as an analytical method, is based on the absorption/reflection of NIR radiation at a specific region or wavelength by the samples of various natural products. While it works fairly well in many cases, it is often difficult to accurately determine what wavelength(s) or region(s) in the near-infrared spectrum carried the most quantitative information about the contents of natural compounds analyzed. In this paper, the assignment of main absorption bands in the spectrum to various probable functional groups, as described above, was done according to literature complied by Workman and Weyer [40] which showed a good agreement with the information for the functional groups in the spectrum given by WinISI software. In the literature, the chemical interpretation for absorption/reflection of NIR radiation at a specific wavelength often varies according to what experimental materials and chemical components were treated for NIR analysis [32] , [41] and [42] . However, the NIRS technique has successfully been employed for determining the contents of various natural compounds in food, feed, biomass, and other natural products even without pinpointing chemical information regarding prominent functional groups related to the near-infrared spectrum [8] [17] [21] [33] [36] [41] [43] .
3.2. Laboratory Reference Data for Various Forage Quality Attributes
The descriptive statistics including, range, mean, median, standard deviation (SD), first & third quartiles (Q1 and Q3), and inter-quartile distance (IQ) for DM, CP, ADF, NDF, DDM, DMI and RFV of the wildrye sample sets used in the calibration and validation sets are shown in Table 1. Mean values various forage quality attributes in the calibration and validation sets were similar. These were 93.74 versus 93.72% for DM, 12.87 versus 13.04% for CP, 33.60 versus 32.16% for ADF, 53.60 versus 53.43% for NDF, 62.72 versus 63.17 for DDM, 2.30 versus 2.29% (LBW) for DMI, and 108.7 versus 109.9 for RFV in the calibration and validation sets, respectively. Likewise, the median, SD, Q1, Q3, and IQ of the validation sample set were more or less similar to those of the calibration sample set in most cases. In addition, the observed results are in agreement with the results reported by other researchers for the southern forages from grass family [44] .
![]()
Table 1. Descriptive statistics for dry-matter (DM), crude protein (CP), acid detergent fiber (ADF), and neutral detergent fiber contents (NDF) in the Southeastern Wild Rye (Elymus glabriflorus) samples used in both calibration and validation.
aQ1, first quartile; bQ3, third quartile; cSD, standard deviation of mean. IQ, inter-quartile distance (IQ = Q3 − Q1).
3.3. Calibration Models
The calibrations and cross validations statistics of the developed NIRS models for DM, CP, ADF, NDF, DDM, DMI and RFV of wildrye are shown in Table 2. The MPLS regression models developed based on the spectral information from full spectra covering 400 to 2498 nm wavelength (both visible and NIR range) using the second-derivative transformation with scatter correction (by SNVD) of raw reflectance spectra yielded the equations for all seven parameters with relatively higher R2 (>0.8) and lower SEC in the calibrations along with higher 1-VR (>0.8) and lower SECV values in the cross-validations (Table 2) than other regression methods (the actual outcomes not shown) tested in this study. We also observed that the use of this whole visible-NIR range (400 - 2498 nm) resulted in much higher R2 and 1-VR and lower SEC and SECV than when using either just the visible range (400 - 1100 nm) or just the near-infrared range (1100 - 2498 nm) (data not shown). Optimum wavelengths for NIR analysis mostly rely on empirical calibrations for predicting qualitative constituents in agricultural products. This is because of the broad array of chemical compounds present in the samples, which lead to overlapping and perturbed NIR absorption bands [32] .
For all seven parameters, the equations using mathematical treatment 2, 4, 4, 1 were selected over other mathematical treatments because they provided better values of several calibration statistics used as the judging criteria. The selected equations for DM, CP, ADF, NDF, DDM, DMI, and RFV resulted in high values of R2 (0.904, 0.974, 0.896, 0.934, 0.895, 0.924, and 0.932, respectively) as well as 1-VR (0.844, 0.963, 0.820, 0.843, 0.820, 0.823, 0.884, respectively). In addition to R2, the 1-VR is an important statistic in judging the quality of calibration equations. A calibration equation with high R2 may have low 1-VR in cross-validation, which suggests a failure of the model in cross validation and hence not a strong model at all. All seven selected equations in this study were
![]()
Table 2. Near infra-red spectroscopic equation development statistics using MPLS and scatter correction for the NIRS prediction of dry-matter (DM), crude protein (CP), acid detergent fiber (ADF), and neutral detergent fiber contents (NDF) in the Southeastern Wild Rye (Elymus glabriflorus) samples.
aSamples used to develop the model; bNumber of PLS loading factors in the regression model MPLS (modified partial least-squares); cSEC, standard error of calibration; dR2, coefficient of determination of calibration; eSECV, standard error of cross-validation; f1-VR, one minus the ratio of unexplained variance divided by variance or coefficient of determination in cross validation; gRSCD, SD/SECV, the ratio of standard error of cross validation to deviation (SD, standard deviation of reference data in calibration set); hRSCIQ, IQ/SECV, the ratio of standard error of cross validation to inter-quartile distance (IQ, inter-quartile distance in reference data in the calibration set); iLBW, live body weight of the animal.
associated with not only high R2 values, but also with invariably high 1-VR values, suggesting good correlations between reference values and NIRS predicted values in each case. The quality of the equations was also judged based on their RSCD (SD/SECV) values, a widely used selection criterion of the NIR calibration equations ( [34] and [38] ). The RSCD values for the equations each of DM, CP, ADF, NDF, DDM, DMI, and RFV (2.54, 5.16, 2.35, 2.53, 2.36, 2.40, and 2.94, respectively) were all greater than 2.0, indicating a close relationship between reference values and NIRS predicted values [32] , [34] and [38] . The RSCIQ (IQ/SECV) is another such criterion, which has been claimed to be a more robust than RSCD because it is based on inter-quartile distance instead of SD, which better represents the spread of the population [37] . The calculated values of RSCIQ for the selected equations of DM, CP, ADF, NDF, DDM, DMI, and RFV (4.65, 5.92, 1.28, 3.38, 1.35, 2.53, and 2.81, respectively) were also invariably greater than 2.0 for all but ADF and DDM, reconfirming the accuracy of at least for equations developed for DM, CP, NDF, DMI, and RFV. However, the original paper of Bellon-Maurel et al. [37] , where the RSCIQ was proposed as a judging criterion of NIRS calibration model, did not discuss the interpretation of the situation with a low RSIQ, but high RSCD as observed for the ADF and DDM calibrations of this study. Therefore, the acceptance or rejection of a calibration model solely based on RSIQ value (>2.0 or <2.0) remains questionable. The other judging criteria must be taken into due consideration. Based on this trend, the models for ADF and DDM may leave rooms for further improvement, but should not be considered as the failed ones at all, because they yielded acceptable values of all other statistics used in numerous reports as the judging criteria for NIRS calibration models. The independent validation performance merits closely monitoring in such case and could be used as the judging criteria with more emphasis.
Figure 3 shows the plots of laboratory reference values versus NIR predicted values for DM, CP, ADF, and NDF contents for the calibration set. Such plots for DDM, DMI, and RFV are not shown because these are calculated from ADF and NDF contents. The diagonal dashed line in each plot is the 1:1 line. The closeness of the plotted data that points to this line indicates the closeness between the NIR predicted values and the corresponding laboratory reference values. As revealed in the plots for ADF, the values deviate from normal to some extent with higher frequency of the lower values. Such a high density of low values often results in more favorable coefficients of determination than when the values are more evenly distributed over the range (for example with DM, CP, and NDF).
3.4. External Validation
The predictability of the seven NIRS calibration models for DM, CP, ADF, NDF, DDM, DMI, and RFV wildrye developed in this study was tested through external validation with 57 samples. The statistics of such external validation exercise such as r2, bias, bias (limit) (maximum allowable bias), SEP, SEPc (the bias-corrected SEP), SEPc (limit) (the maximum allowable SEPc), slope, RPD (SD/SEPc) and RPIQ (IQ/SEPc) values are presented in Table 3. These statistics were used to evaluate the predictability or
![]()
Figure 3. Scatter plots of NIRS predicted values versus laboratory reference values for the calibration sets. SEC, standard error of calibration. R2, coefficient of determination of calibration. 1-VR, one minus the ratio of unexplained variance divided by variance, also called coefficient of determination in cross validation. SECV, standard error of cross-validation.
reliability of the calibration models. An NIRS calibration model is considered robust and reliable when it can produce lower bias [(lower than the bias (limit)] and SEPc [(lower than the SEPc (limit)] with r2 and slope close to 1.0, and RPD and RPIQ values greater than 2.0 in external validation with samples independent of calibration sample set [32] [34] [37] [38] . Based on these criteria, the r2, and RPD values obtained during external validation of all seven models were high enough (Table 3) to indicate good agreement between reference values and NIRS predicted values, similar to the corresponding statistics obtained during the calibration development (Table 2). The r2 values were greater than 0.8 for all parameters. Likewise, the RPD values were invariably greater than 2.0-threshold value. However, RPIQ values for ADF and DDM were lower than 2.0 threshold value just as observed for RSCIQ discussed earlier. The slope was greater than 0.90 for all seven parameters; it was very close to 1.0 for DM, CP, NDF, and RFV. More than 90% of the samples included in the validation set were successfully predicted by all calibration models except ADF with difference between measured and predicted values smaller than three times original SECV. For ADF model, this was 86%. Figure 4 depicts the plots of NIRS predicted values versus laboratory reference values in the validation set for DM, CP, ADF, and NDF contents, which shows the significant relationship between NIRS and reference.
![]()
![]()
Figure 4. Scatter plots of NIRS predicted values versus laboratory reference values for the validation sets. n, samples (independent) used to monitor the model. r2, coefficient of determination in external validation. SEP, standard error of prediction. SEPc, the bias-corrected standard error of prediction.
4. Conclusion
All seven NIRS calibration models presented and discussed in this paper showed accurate prediction abilities for some important forage quality parameters of wildrye related to animal nutrition. Thus, these models can be reliably applied in the routine analysis of these properties. Once the samples are dried and ground, this nondestructive NIRS method is very rapid and could simplify the analysis of qualitative factors of interest by replacing tedious laboratory methods. Despite the complicated nature of spectral information and somewhat speculative interpretation of the absorption/reflection bonds, the well-researched and widely used chemometric techniques are able to establish valid quantitative relationship between the spectral information and the parameters measured for the samples leading to an acceptable tool to be used for predicting the parameters in unknown samples. As a result, properly developed and validated NIRS calibration models, as presented in this study, become a very reliable method for predicting unknown samples that have resemblance with the samples utilized in the calibration development. When high throughput in analysis is demanded by the industry, such as in integrated forage based livestock production industries, NIRS method is especially suitable because it replaces the time consuming laboratory methods without sacrificing accuracy and precision. Thus, it consistently plays its role as a reliable analytical tool for decision allowing determination of multiple values (e.g. DM, CP, ADF, NDF etc.) in a single analytical procedure thereby assisting in timely decision making on strategic use of nutritional supplements or adjustments in ration formulation to efficiently sustain milk, meat, or fiber production. Although development of an NIRS laboratory entails significant initial start-up costs, it is relatively inexpensive in the long term. It is also considered as “cheap” and “green chemistry” because it does not involve any chemicals and does not generate any hazardous wastes. Despite many advantages of NIRS method, its prediction becomes merely an unreliable extrapolation when it is improperly used or abused such as applying on predicting samples that are very different from the calibration sample set. This can happen due to a variety of reasons such as differences in species, cultivation area, drying and storage methods, influences of genetic and environmental variation, and others. However, such problem of spectral outliers should be watched out and solved by updating, expanding, and improving the initially developed and validated calibrations (such as the ones presented here) by including future samples from different environments and species and covering a wider range of the parameters to impart further robustness to the current calibration models. This should be a routine practice of the NIRS laboratories.
Acknowledgements
This publication is a contribution of the “Mississippi Agricultural and Forestry Experiment Station”. The authors are grateful for the Agricultural and Environmental Services Laboratories, University of Georgia, for hosting NIRS calibration development and validation activities. This research did not receive any specific grant from funding agencies in the public, commercial, or not-for-profit sectors.
Abbreviations
ADF, Acid Detergent Fiber;
CP, Crude Protein;
DDM, Digestible Dry Matter;
DMI, Dry Matter Intake;
NDF, Neutral Detergent Fiber;
NIRS, Near-infrared Reflectance Spectroscopy;
RFV, Relative Feed Value;
RPD, Ratio of Performance to Deviation;
RPIQ, Ratio of Performance to Inter-Quartile distance;
RSCD, Ratio of Standard Error of Cross Validation to Deviation;
RSCIQ, Ratio of Standard Error of Cross Validation to Inter-Quartile Distance.