1. Introduction
Inorganic polyphosphate (PolyP) is multifunctional biopolymer performing many cellular functions in all living cells, from prokaryotic to human [1] - [8] . The ability of PolyP metabolizing enzymes to catalyze the conversion of other substrates is one of the causes of the involvement of these proteins in regulatory pathways. For example, the gppA exopolyphosphatase splits Pi from guanosine 5’-triphosphate, 3’-diphosphate and guanosine 5’-diphosphate, 3’-diphosphate, bacterial second messengers [9] . Some bacterial exopolyphosphatases display NTPase activities [10] . The enzymes belonging to the polyphosphate kinase 2 subfamily catalyze nucleoside monophosphate phosphorylation [11] . The exopolyphosphatase of Pseudomonas aeruginosa is a polyphosphate: ADP phosphotransferase [12] . The Saccharomyces cerevisiae protein DDP1 (a diadenosine and diphosphoinositol polyphosphate phosphohydrolase) possess an endopolyphosphatase activity [13] . The exopolyphosphatase PPX1 of S. cerevisiae hydrolyzes adenosine-tetraphosphate phosphohydrolase and guanosine-tetraphosphate phos- phorhydrolase activities [14] . The polyphosphatase PPN1
(http://www.uniprot.org/uniprot/Q04119) of S. cerevisiae degrades PolyP both by clea- ving Pi from the chain end and by fragmenting long-chain polymers into shorter ones [15] [16] . Pi release was predominant in the presence of Co2+, while the fragmentation of high-molecular PolyP was predominant in the presence of Mg2+ [16] . No other substrates of this enzyme are known; the pyrophosphatase activity of PPN1 was extremely low [17] . We have attempted to use the pure recombinant PPN1 [18] to reveal PolyP in the PCR reaction products with the high level of pyrophosphate [19] . In these experiments, the treatment of dNTP mixture with PPN1 resulted in Pi release; i.e., PPN1 catalyzed the reaction:
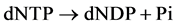
This study was aimed at characterizing the dNTP phosphohydrolase activity of PPN1 with dATP as a substrate.
2. Materials and Methods
2.1. Strain Growth
The ΔPPN1 mutant strain CRN of S. cerevisiae was obtained from N. Rao and A. Kornberg [15] . The strain CRN/pMB1_PPN1 Sc of S. cerevisiae overexpressing the polyphosphatase PPN1 was designed by Eldarov and co-authors earlier [20] . Yeast strains were grown in a synthetic minimal YNB medium containing (per 1 l) 1.7 g of bacto yeast nitrogen bases (Difсo, Detroit, USA), 20 g of glucose, by 20 mg of L-tryptophane, L-histidine, L-methionine and adenine, and 60 mg of L-leucine. Uracyl (20 mg/l) was added for cultivation of the strain CRN. The cells were cultivated up to the stationary growth stage under shaking (145 rpm/h) at 29˚C.
2.2. PPN1 Purification
The cellular extracts were obtained as described [18] . The cellular extract of the strain CRN/pMB1_PPN1 Sc was used for purification of polyphosphatase PPN1 by a combination of the methods designed earlier for obtaining pure PPN1 [18] . The cellular extract was supplemented by ammonium sulfate up to 50% saturation and incubated for 1 h, followed by centrifugation at 12,000 g for 20 min. The supernatant was absorbed on the Butyl-Toyopearl 650 M equilibrated with 50 mM Tris-HCl, pH 7.2, with 50% ammonium sulfate. In 45 min, the resin was precipitated at 4000 g for 3 min and washed three times with 50 mM Tris-HCl, pH 7.2, containing 50% ammonium sulfate. For polyphosphatase elution, the resin was washed four times with 50 mM Tris-HCl, pH 7.2, containing 25% ammonium sulfate. Triton X-100 (0.05%) was added to the preparation. After ultrafiltration through an YM-10 membrane, the solution was applied to a DEAE-Toyopearl 650 M column (1.5 × 7.5 cm) equilibrated with 25 mM Tris-HCl, pH 7.2, with 0.1% Triton X-100. The column was washed with 50 ml of the same buffer and then with 50 ml of 0.1 M KCl in the same buffer. The polyphosphatase was eluted at a flow rate of 24 ml/h with an increasing KCl concentration (0.1 - 0.8 M) in the same buffer. The gradient volume was 200 ml. The fractions with the polyphosphatase activity were pooled, concentrated by ultrafiltration, and incubated with heparin-agarose for 2 h. The resin was pre-equilibrated with 25 mM Tris-HCl, pH 7.2, with 0.1% Triton X-100. The heparin-agarose washed with 10 ml of the buffer and then 5-fold with 5 ml of 0.5 M KCl in the same buffer, using centrifugation at 1500 g for 5 min. Then the resin was washed twice with 5 ml of 0.7 M KCl in the buffer; the exopolyphosphatase was desorbed with 5 ml (4-fold) of 1 M KCl in the same buffer. The preparation had a specific activity with polyP 208 - 2000 E/mg of protein. The protein was assayed with Pierce Bradford reagent after precipitation with trichloroacetic acid. The preparation was stored at −20˚C.
2.3. Enzyme Activities Assay
The enzyme activities were assayed at 30˚ in 0.1 ml of 50 mM Tris-HCl, pH 7.2, containing 0.1 mM CoSO4 and 200 mM NH4Cl. The concentrations of MgSO4, MnSO4, ZnSO4, and other additives are indicated in the legends to the tables and figures. The amount of the enzyme releasing 1 nmole of phosphate (Pi) per 1 min was taken as a unit of enzyme activity (mU). Inorganic polyphosphate with an average chain length of 208 phosphate residues (polyP208) (Monsanto, USA) was used as a substrate of exopolyphosphatase at the concentration of 2.5 mM (the concentration was estimated by phosphorus). PolyP208 was purified from pyrophosphate and orthophosphate as described [21] . The dATP (GE Healthcare Bio-Sciences Corp, Lithuania), ATP, ADP and pyrophosphate (Sigma) were used at the concentration of 2 mM. Heparin was from Spofa, Czech Republic. The released Pi was assayed with malachite green [22] with an immunoplates spectrometer (Sapfir, Russia). The Pi content in the samples without the enzyme was assayed as a control.
2.4. Statistics
The assays were performed in triplicate; the mean values and standard deviations were calculated by Excel. The correlation coefficient was calculated using
http://www.alcula.com/calculators/statistics/correlation-coefficient/.
3. Results
The purified PPN1 catalyzed the release of Pi from dATP (Table 1). The other exopolyphosphatase of S. cerevisiae, PPX1 (kindly provided by L. Lichko) [23] , did not split Pi from dATP (Table 1). It is surprising, because PPX1 is more active with polyP3 [23] and with adenosine-tetraphosphate and guanosine-tetraphosphate [14] . The PPN1 activity was maximal with long-chain PolyP and weak with PolyP3 [17] [18] . The dATP phosphohydrolase activity of pure PPN1 was ~7-fold lower compared to the exopolyphosphatase activity (Table 1). The activity of PPN1 with ATP was nearly twofold lower than with dATP; the activity with ADP and PPi was still lower (Table 2). The concentration dependence of dATPase activity of this enzyme corresponded to the Michaelis-Menten equation, and Km was 0.88 ± 0.14 mM.
The exopolyphosphatase activity of PPN1 displays the non-Michaelis kinetics, with the apparent Km of 0.0035 and 1.1 mM with PolyP208 and PolyP3, respectively [17] . The substrate affinity and hydrolysis rate of PolyP3 [17] and dATP were similar. Both exopolyphosphatase and dATPase activities of PPN1 were stimulated by NH4Cl (200 mM) nearly twofold (not shown).
Figure 1 shows the dependence of dATPase activity on the concentration of divalent cations. Co2+ was more effective than Mn2+; Mg2+ and Zn2+ had no stimulatory effects. Heparin and PPi inhibited this activity and the inhibitory effect of ADP was low (Figu- re 2). The effects of divalent cations and the above inhibitors on the dATPase (Figure 1, Figure 2) and exopolyphosphatase [18] activities of PPN1 were similar.
The dATP phosphohydrolase (dATPase) activity of cellular extract of the PPN1- overexpressing yeast strain was several-fold higher than that in the parent strain (Table 3). This increase was comparable with the increase in exopolyphosphatase activity. In addition, the dATPase and exopolyphosphatase activities of the cellular extract of trans- formant cells were similarly inhibited by heparin, the known suppressor of polyphosphatases [1] , while the dATPase activity of the cellular extract of ΔPPN1 mutant was little affected by heparin. Probably, some other enzymes perform the dATPase activity in this strain.
![]()
Table 1. The exopolyphosphatase and dATP phosphohydrolase activities of pure PPN1 and PPX1 of S. cerevisiae.
![]()
Table 2. Pi release by PPN1 from some substrates (2 mM).
![]()
Figure 1. The effects of Co2+ (white squares), Mg2+ (open circles), Zn2+ (black triangles), and Mn2+ (open triangles) on the dATP phosphohydrolase activity of polyphosphatase.
![]()
Figure 2. The effects of inhibitors on the dATP phosphohydrolase activity of polyphosphatase PPN1. The activity was assayed in 50 mM Tris-HCl, pH 7.2, containing 0.1 mM CoSO4 and 200 mM NH4Cl.
![]()
Table 3. The hydrolysis of PolyP208 and dATP by the cellular extracts of ΔPPN1 and PPN1- overexpressing strains of S. cerevisiae.
4. Discussion
In this study, we have found the dATP phosphohydrolase activity of the S. cerevisiae protein РРN1. The activity is similar to the exopolyphosphatase activity of this protein in the dependence on divalent cations, stimulation by NH4 ions, and inhibition by heparin and PPi. The activity decreased in the following order: dATP > ATP > ADP. The preliminary data on the hydrolysis of dNTP mixture suggest the ability of hydrolysis of other dNTP. It should be noted that the PPX1 exopolyphosphatase of S. cerevisiae hydrolyzes neither dATP (this study) nor dNTP mixture [19] .
The catabolism of dNTPs is performed by many enzymes. First, the alkaline and acid phosphatases are able to hydrolyze these substrates (http://www.brenda-enzymes.org).
Second, the DEAH-box splicing factor Prp22 of yeasts can hydrolyze all common NTPs and dNTPs with a comparable efficiency [24] , catalyzing the reaction:
(d) nucleoside triphosphate + H2O → (d) nucleoside diphosphate + phosphate (EC 3.6.1.15). Finally, deoxynucleotide triphosphate triphosphohydrolases (dNTPases) hydrolyze deoxynucleotide triphosphates (dNTPs) into nucleosides and tripolyphosphate (3.1.5.B1, http://www.brenda-enzymes.org). In mammalian cells, the sterile alpha motif and HD domain-containing protein 1 (SAMHD1) is a dNTPase and a major regulator of cellular dNTP levels [25] . This protein prevents the infection of nondividing cells by retroviruses, including HIV, by depleting the cellular dNTP pool [25] . The importance of dNTP metabolism and SAMHD1 in cancer development was discussed [26] . It seems that the dNTP catabolism becomes particularly important under the conditions when a decrease in proliferative activity is necessary for the survival of an organism or a cell population. For further investigation of the role of PPN1 as a putative regulator of cellular dNTP level in yeast cells, it is necessary to take into account the following two facts. First, PPN1 gene is responsible for the exopolyphosphatase activity in yeast nuclei [27] . Second, ΔPPN1 mutants displayed the impairment of the cell cycle when the cells were grown under Pi limitation [4] .
5. Conclusion
Yeast polyphosphatase PPN1 is deoxyadenosine triphosphate phosphohydrolase while the exopolyphosphatase PPX1 did not split Pi from dATP. The exopolyphosphatase and deoxyadenosine triphosphate phosphohydrolase activities of this protein are similar in ion and inhibitor effects.
Acknowledgements
The authors thank Elena Makeeva for her help with preparing the manuscript. This study is supported by Russian Basic Research Foundation (Grant 14-04-00515).
List of Abbreviations
PolyP―inorganic polyphosphates, PolyP208―inorganic polyphosphates with an average chain length of 208 phosphate residues, PolyP3―tripolyphosphate, Pi―ortho- phosphate, PPi―pyrophosphate; dATPase―deoxyadenosine triphosphate phosphohydrolase.