Rubber Seed Kernel as Potent Solid Substrate for the Production of Lipase by Pseudomonas aeruginosa Strain BUP2 ()
1. Introduction
Being a versatile biocatalyst, microbial lipases (triacylglycerol acylhydrolase, EC 3.1.1.3) offer great potentials in biological as well as industrial applications [1] . Though industry mostly prefers submerged fermentation (SmF) strategy for the production of lipase, solid-state fermentation (SSF) received more attention recently due to its cost effectiveness and higher productivity. However, numerous studies attempted for the production of lipase on solid substrate, in which different solid agricultural residues such as cakes from deoiled coconut and groundnut kernels; husks of rice, lentil and wheat; residue from banana, melon, soybean and watermelon; and brans from wheat and rice have been used [2] -[4] . Oil cake is a cheap byproduct emerged out of oil extraction, which is mainly used in animal and chicken feeds. In this connection, large potential of rubber seed (oil and cake) as substrate for the cultivation of miocroorganisms is neglected.
India is one of the largest producers of rubber in the world, and every year huge quantity of rubber seed is treated as agricultural waste. Rubber seed oil is rich in fatty acids: i.e., 39.6% linoleic acid, 23.5% oleic acid, 19.3% linolenic acid, 8.7% stearic acid 2% palmitic acid, apart from carbohydrate (24.21%) and protein (22.17%). Owing to the high nutritional content of the agro-industrial residues, oil cakes are considered as valuable solid substrates for the production of enzymes by growing suitable microorganisms on it under water-re- stricted environment, the SSF. Moreover, usage of agricultural residues with an industrial perspective is one of the best strategies for the better agricultural waste management and abatement of environmental pollution problem due to agricultural residues.
Though many microorganism such as bacteria, yeast, actinomycetes and fungi shown to have the potential for the production of lipase, species of genera Bacillus, Pseudomonas, Staphylococcus, Candida, Geotrichum, Aspergillus, Mucor, Penicillium, Rhizopus and Rhizomucor are considered as the best producers of lipase [5] . Pseudomonas aeruginosa strain BUP2 (MTCC No. 5924), a new bacterial strain reported from this laboratory [6] was used in this study. Thus, the present study investigates the efficiency of the ground kernels of rubber seed, coconut and groundnut or deoiled cakes as solid substrate for the production of lipase by P. aeruginosa strain BUP2.
2. Materials and Methods
2.1. Bacterial Culture
Pseudomonas aeruginosa strain BUP2 (MTCC No.5924), a new bacterial strain already reported from this laboratory was used throughout the study, which was isolated from the rumen of Malabari goat [6] .
2.2. Chemicals
Analytical grade chemicals from Hi Media Laboratories (Mumbai, India) were used for the study. The p-nitro phenyl palmitate (pNPP), substrate for lipase assay was purchased from Sigma Chemical Co., USA.
2.3. Medium and Inoculum
P. aeruginosa BUP2 was maintained 4˚C on mineral salt-oil-agar medium. Seed culture (12 h old) was prepared in the BUP medium by inoculating a loopful of stock culture of bacterium into it [6] , which was incubated at 37˚C and 130 rpm.
2.4. Medium for SSF
For the cultivation of P. aeruginosa strain BUP2, kernels of 3 types of nuts/seeds (coconut, groundnut and rubber seed) were used in the ground form or as their deoiled cake for this study. Before grinding, the kernel was chopped into small pieces and dried in an oven for 24 h at 60˚C. The coconuts and groundnuts were purchased from local market, while rubber seeds were collected from a local plantation. The deoiled cakes were prepared after extracting oil using an expeller manually.
2.5. Solid-State Fermentation (SSF) Strategy
Two grams of ground kernel or their cakes was moistened with 10 ml of BUP medium. All preparations in the flask were autoclaved at 121˚C for 15 min, and inoculated with 0.1 ml of inoculum (seed culture) under aseptic condition; and the inoculated media were incubated at respective conditions. In order to check lipase production, fermented samples were assayed at regular intervals of 24 h for 4 days.
2.6. Effect of pH
The solid substrate (ground seeds/cake) was moisturised with BUP medium having varying pH concentration (4, 5, 6, 7 and 8) was inoculated with P. aeruginosa strain BUP2 and incubated at 25˚C. At regular intervals of 24 h, the fermented matter was analyzed for estimating the production of lipase.
2.7. Effect of Temperature
To estimate the role of different temperature (25˚C, 28˚C, 30˚C, 32˚C) on lipase production, the ground kernel or cake was inoculated with P. aeruginosa strain BUP2 after moisturising it with BUP medium having optimum pH, and assayed for lipase activity at regular intervals of 24 h.
2.8. Effects of Substrate Concentration
Effect of substrate concentration [(10%, 20%, 30%, 40% and 50% (w/v)] on lipase production was checked under the optimized conditions of pH and temperature. The fermented samples were regularly withdrawn at every 24 h interval, and assayed for lipase activity.
2.9. Lipase Extraction
Lipase was extracted from the fermented solid substrate by the method of [7] . Crude lipase was extracted by mixing 1 g of fermented substrate with 5 ml of 0.1 M Tris-HCl buffer (pH 8.0), and mixed well on a vertex mixer. After centrifugation (8000 × g for 10 min, 5˚C), the supernatant was collected for lipase (crude) assay.
2.10. Lipase Assay
Lipase activity in the cell and debris free supernatant was determined as described by [8] . The reaction mixture (1.8 ml) contained 0.15 M NaCl and 0.5% triton X-100 in 0.1 M Tris-HCl buffer (pH 8.0) and 200 µl supernatant (or double distilled water in control), which was pre-incubated (10 min) at 37˚C in a water bath. Subsequently, 20 µl pNPP (p-nitrophenylpalmitate) in 50 mM acetonitrile was added, and incubated further at 37˚C for 30 min. The quantity of liberated p-nitrophenol was recorded at λ405. One unit of lipase activity is defined as the quantity of enzyme required for liberating 1 µmol of p-nitrophenol under the standard assay conditions. Lipase activity was calculated using the following formula:
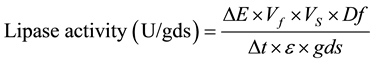
where ΔE is the absorbance at 405 nm; Vf is the final volume of reaction mixture; VS is the volume of crude supernatant (lipase) used; Df is the dilution factor (i.e., total extracted volume from 1g fermented matter); Δt is the time of incubation in min; ε is the extinction coefficient (0.017); gds is the dry weight in grams (i.e., dry matter of the 1 g fermented matter used for extracting lipase).
3. Statistics
All studies were repeated at least thrice, and an average of 3 values is presented with standard deviation. Microsoft Excel was used to draw the figures.
4. Results
4.1. Effect of pH
The effect of pH on lipase production was investigated at different pH using different ground kernels or their cakes. It showed that slightly acidic or neutral pH supported the maximum production of lipase. Among the different substrates used, rubber seed flour supported the maximum production of lipase (340 U/gds) at pH 6, followed by groundnut flour and its cake (181 and 168 U/gds, respectively) at pH 7; whereas coconut oil flour and its cake supported lipase production at pH 6, but in lesser quantities (130 and 103 U/gds, respectively) (Figure 1).
4.2. Effect of Culture Temperature
The effect of temperature on the production of lipase by P. aeruginosa strain BUP2 was determined at various temperature (25˚C to 32˚C); of which, all the substrates supported the maximum production of lipase at temperature range between 28˚C and 30˚C. Rubber seed flour, coconut flour and groundnut cake supported the maximum lipase production (871, 174 and 203 U/gds, respectively) at 28˚C; while coconut cake (344 U/gds) and groundnut four (397 U/gds) supported the maximum production of lipase at slightly higher temperature (30˚C) (Figure 2).
4.3. Effect of Substrate Concentration on Lipase Production
Five solid substrates (coconut flour, coconut cake, groundnut flour, groundnut cake and rubber seed flour)
![]()
Figure 1. Effect of pH on lipase production by P. aeruginosa strain BUP2 on various flours (COP―coconut powder, GOP―groundnut powder, RSP―rubber seed powder) and cakes (COC― coconut cake, GOC―grondnut cake) moisturised with BUP medium.
![]()
Figure 2. Effect of temperature on lipase production by P. aeruginosa strain BUP2 on various flours and cakes moisturised with BUP medium.
enriched with BUP medium were tested for their effect on lipase production by P. aeruginosa strain BUP2. Of them, the maximum lipase production (871 U/gds) was supported by 20% (w/v) of rubber seed flour under optimized condition (pH 6, 28˚C and 48 h incubation) (Figure 3); and 20% (w/v) of groundnut flour (pH7, 30˚C and 48h incubation) supported the production of 398 U/gds lipase (Figure 4). Coconut cake (20%, w/v) and coconut flour (40%, w/v) supported 327 and 311 U/gds lipase production, respectively (Figure 5 & Figure 6). Groundnut cake (30%, w/v) supported comparatively lesser lipase production (244 U/gds), which was at 72 h of incubation (Figure 7).
5. Discussion
The primary objective of this work was to investigate the suitability of agricultural product-based fermentation medium for lipase production by P. aeruginosa strain BUP2 (MTCC No 5924). Large quantities of deoiled kernels of seeds/nuts are generated as by-product during the extraction vegetable oil, which would fetch low price to the farmers. Rubber seed (oil or kernel) is not considered as a healthy source of cooking oil or its industrial
![]()
Figure 3. Lipase production profile of P. aeruginosa strain BUP2 on 10, 20, 30, 40 or 50% (W/v) of rubber seed flour enriched with BUP medium.
![]()
Figure 4. Lipase production profile of P. aeruginosa strain BUP2 on 10, 20, 30, 40 or 50% (w/v) of groundnut flour enriched with BUP medium.
![]()
Figure 5. Lipase production profile of P. aeruginosa strain BUP2 on 10, 20, 30, 40 or 50% (w/v) of rubber seed coconut cake enriched with BUP medium.
![]()
Figure 6. Lipase production profile of P. aeruginosa strain BUP2 on 10, 20, 30, 40 or 50% (W/v) of coconut flour enriched with BUP medium.
uses are limited. Thus, a few of them were explored in this study for their utility as substrate for the production of lipase, thereby increasing their market value. SSF is mainly employed with fungi for the production of extracellular enzymes such as lipase, cellulase, amylases alkaline proteases and xylanase. In general, bacterial cultures are not considered suitable for performing SSF due to the higher water activity requirement for their growth. However, numerous studies showed that bacterial cultures can well be adapted or it can be manipulated for SSF processes [9] -[11] . Bacteria can efficiently produce α-amylase, alkaline protease and inulinase by species of Bacillus [12] [13] , Pseudomonas and Staphylococcus [9] -[15] .
In the light of several advantages of SSF, lipase production from the strain of P. aeruginosa strain BUP2 was attempted on agricultural products. The rubber seed flour supported the maximum production of lipase. To the best of our knowledge, this is the first report showing rubber seed flour as a potent substrate for the production of lipase via SSF. A few studies reported oil cakes as the solid substrate-cum-inducer for the production of lipase by SSF [8] [16] [17] . The utility of mixed-solid substrate contains wheat bran and coconut oil cake for lipase
![]()
Figure 7. Lipase production profile of P. aeruginosa strain BUP2 on 10, 20, 30, 40 or 50% (w/v) of groundnut cake enriched with BUP medium.
production by Candida rugosa [10] . Recently, efficacy of Pseudomonas sp. strain BUP6 for the production of lipase (107 U/gds) on groundnut cake was illustrated by [18] . Deoiled cake from Jatropha seed was used as a support for the production of lipase (1084 U/g) from P. aeruginosa PseA through SSF [19] .
6. Conclusion
This study showed the utility of agricultural products, especially cheaply available rubber seed as solid medium for the production of valuable lipase. This would enable the rubber farmers to get additional income from their agriculture. Moreover, the ability of P. aeruginosa strain BUP2 to grow on rubber seed medium and secretion of extracellular lipase are highly promising. Moreover, elimination of limarin, a known toxin from the rubber seed by fermentation, is an added advantage, showing its utility in cattle and poultry feeds. In this context, efficiency of P. aeruginosa strain BUP2 to grow on other flours and cakes and secretion of lipase are yet other promising signs of utility of this novel bacterium.
Acknowledgements
The financial assistance (Grant No. 026/SRSLS/2012/CSTE) from Kerala State Council for Science, Technology and Environment (KSCSTE), Government of Kerala, is gratefully acknowledged.