Visualization of Pareto Solutions by Spherical Self-Organizing Map and It’s acceleration on a GPU ()
1. Introduction
With the rapid rise of hardware and software performance in recent years, computer-aided numerical simulations are being used more and more for designing products. These include construction drawings with CAD (Computer-Aided Design), optimizations of design parameters, and validations of candidate solutions, among others.
It is often effective to formalize problems as optimization problems in order to employ numerical simulations as design aids. An optimization problem is one in which design variables are determined so that one or more objective functions yield the highest or lowest possible values under certain conditions. Real-world optimization problems tend to have multiple objective functions that pose trade-offs among each other: this type of problem is called a multiple-objective optimization problem. Because of the trade-offs, a multiple-objective optimization problem has multiple optimal points, each of which is the most fit for a certain objective function. The set of these solutions is called the Pareto-optimal solution set. Many approaches have been proposed on how to reach the Pareto-optimal solution set [1].
In product design, an important task is to choose the best solutions from a Pareto-optimal solution set. It is required, then, to visualize the solution set and show the designer what characteristics each of the solutions have in relation to other solutions and in relation to the containing set. However, whereas each solution in a multipleobjective optimization problem is a multi dimensional value consisting of many objective function values and design variables, the human eye can distinguish and comprehend merely three dimensions at most. This gap needs to be bridged in order to present an intuitive visualization.
One attempt at this challenge is the self-organizing map (SOM) [2], a kind of artificial neural network. A SOM is a means of translating high-dimension data to low-dimension data and is useful for visualizing a Paretooptimal solution set, typically in three or less dimensions.
Meanwhile, practical use of data analysis on the job site demands short execution times. Although future improvements in hardware performance will allow larger and larger problems to be computed in realistic timeframes, thus enabling a wider range of design aids, recent years have seen the increase of processor frequency hit the ceiling. Instead, grounds for performance gains are shifting to many-core architectures, namely graphic processing units (GPUs). Many-core architectures have multiple processing cores and yield exceptionally good performance with parallel algorithms. Thus, when handling large problems, one should employ highly parallel algorithms that are computed faster with many-core architectures.
While SOMs require a vast amount of computations, they also have high loop-level parallelism and can be expected to perform well on many core architectures [3].
Based on these premises, we explored the visualization of Pareto-optimal solutions using SOMs (spherical SOMs [4] specifically) and evaluated the system using a GPUbased implementations. Spherical SOMs have been reported [5] to be less vulnerable to distortions compared to plane SOMs due to the fact that they have no edges. They also show a comparatively rational spatial relationship among the data points and can be clustered more accurately [4,6].
In this paper, we discuss first the parallelizing of the spherical SOM algorithm, then evaluate its performance with a GPU-based implementation. Subsequently we discuss the visualization of Pareto-optimal solution sets for two optimization problems: test functions commonly used in multi-objective genetic algorithms, and a problem concerning the design of diesel engines, thus pointing out the effectiveness of spherical SOMs and their advantages over planar SOMs.
2. Multi-objective Optimization Problems
A problem where multiple objectives must be optimized at the same time is called a multi-objective optimization problem. A multi-objective problem is generally formalized as Equation (1): there are n design variables x, and k objective functions f(x) are maximized (or minimized) under m constraints g(x).
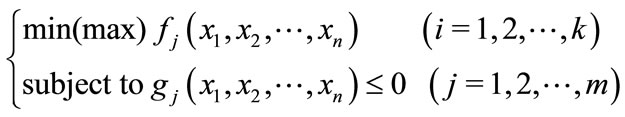 (1)
(1)
Because it is often the case with multiple-objective optimization that there are trade-offs among objective functions, not all objective functions  can be optimized at the same time. Therefore we aim for a set of Paretooptimal solutions rather than a single optimal solution. Pareto-optimal solutions are defined by the solutions’ dominance over each other. Below is the definition of dominant solutions in multiple-objective optimization problems (when all objective functions are optimal when minimized):
can be optimized at the same time. Therefore we aim for a set of Paretooptimal solutions rather than a single optimal solution. Pareto-optimal solutions are defined by the solutions’ dominance over each other. Below is the definition of dominant solutions in multiple-objective optimization problems (when all objective functions are optimal when minimized):
Definition of dominance:
When ( )
)
(a)  dominates
dominates 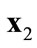 when
when

(b)  strongly dominates
strongly dominates  when
when

A solution  that dominates
that dominates  is better than
is better than  in the general sense. The intent then is to search for nondominated solutions. Based on this definition of dominance, below is the definition of Pareto-optimal solutions:
in the general sense. The intent then is to search for nondominated solutions. Based on this definition of dominance, below is the definition of Pareto-optimal solutions:
Definition of Pareto-optimal solutions:
When (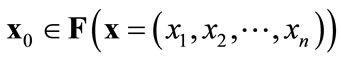 )
)
(a)  is a weak Pareto-optimal solution when there are no solutions
is a weak Pareto-optimal solution when there are no solutions 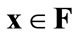 that strongly dominate
that strongly dominate 
(b)  is a Pareto-optimal solution when there are no solutions
is a Pareto-optimal solution when there are no solutions  that dominate
that dominate 
Figure 1 shows an example of Pareto-optimal solutions when there are two objectives (k = 2). Solid dots show Pareto-optimal solutions. Open dots are inferior in every way to Pareto-optimal solutions. Thus we can ignore the white dots and obtain the front consisting of Pareto-optimal solutions, which is shown as the dashed curve (the Pareto-optimal front).
One way to visually investigate what kinds of Paretooptimal solutions constitute a Pareto-optimal solution set is SOM.
3. SOM
3.1. SOM Overview
SOM is a type of neural network trained by unsupervised learning and is known as a means of data mining. SOM can map a group of data into an arbitrary number of dimensions without distorting correlations among these data points. Thus it is useful for visualizing high dimension data into lower dimensions (typically two or three) so that characteristics can be grasped intuitively. It also lets us cluster the solution set and analyze its traits.
3.2. SOM Algorithm
A SOM consists of two layers: the competitive layer and the input layer. The competitive layer is where neurons are placed, and the input layer is the group of input data points. Neurons on the competitive layer are updated as the SOM is trained with successive read-ins from the input layer.
An input data point, defined as , is presented to all neurons in the network. Any given neuron i has a weight vector with number of dimensions n, corresponding to the dimensions of the input data. A neuron’s weight vector for a given time t is defined as
, is presented to all neurons in the network. Any given neuron i has a weight vector with number of dimensions n, corresponding to the dimensions of the input data. A neuron’s weight vector for a given time t is defined as . Learning coefficient α(t), neighborhood function hci(t), and neighborhood range σ(t) are also defined. A basic SOM algorithm is as follows:
. Learning coefficient α(t), neighborhood function hci(t), and neighborhood range σ(t) are also defined. A basic SOM algorithm is as follows:
1) Initialize competitive layer With  and the repetition count
and the repetition count , all neurons
, all neurons  in the competitive layer are set with initial weight vectors.
in the competitive layer are set with initial weight vectors.
2) Compute distances Input data point  is presented to the competition layer and Euclidian distances from the neurons
is presented to the competition layer and Euclidian distances from the neurons
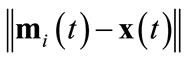 are computed.
are computed.
3) Find the winner neuron The winner neuron c where
 is determined.
is determined.
4) Train The winner neuron c and all neurons within distance dci from it are adjusted by applying Equations (2). The neighborhood function hci(t) is defined by Equations (3) using the learning coefficient α(t). α(t) and the neighborhood range σ(t) are defined by Equations (4) and (5) respectively.
 (2)
(2)
 (3)
(3)
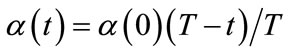 (4)
(4)
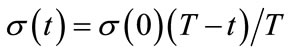 (5)
(5)
5) Terminate If , set
, set  and return to step (2). Otherwise exit the loop.
and return to step (2). Otherwise exit the loop.
4. Spherical SOM
4.1. Spherical SOM Overview
When analyzing a Pareto-optimal solution set, it is important not only to grasp where the solutions are clustered, but also to see what other neighboring solutions exist for a given solution, or how the solutions are positioned in relation to each other.
A map generated with SOM shows similar data points close to each other, which is helpful in assessing the similarities of the solutions. However, the fact that a pair of data points is placed far from each other on the map does not necessarily mean that the similarity is low. A designer analyzing a Pareto-optimal solution set would prefer similar data points to appear as close to each other on the map as possible.
This apparent similarity (or proximity) of the solutions depends heavily on the form of the competitive layer used in SOM. A competitive layer in the shape of a twodimensional plane often yields distortions in proximity because edges exist, and the number of neurons in the neighborhood range differs from neuron to neuron. To avoid this, SOMs with competitive layers in closed forms such as tori or spheres have been proposed.
Torus SOMs are easy to implement as an extension to plane SOMs, but the resulting visualization takes the form of a donut, which is difficult to interpret with human eyes, whereas spherical SOMs place the neurons on the surface of a sphere, making it easier to grasp the spatial relations. Thus spherical SOMs are suitable for visualization.
4.2. Implementation of a Spherical SOM
4.2.1. Data Structure
Neurons in a spherical SOM are commonly placed in a geodesic dome, a type of quasi-regular polyhedron [5]. A geodesic dome is an approximated sphere constructed by halving each edge of a regular polyhedron, thus recursively increasing the number of surfaces. Neurons are placed on the apices of the geodesic dome, forming a hexagonal grid. Apices on a geodesic dome give the neurons a more uniform arrangement on the surface of a sphere compared to other methods such as polar coordinates. In this study we used a regular icosahedron, the regular polyhedron with the largest possible number of surfaces, for the original approximation of the sphere to further ensure uniformity.
A geodesic dome, when unfolded via the edges of the icosahedron, forms a net that can be mapped to and handled in two dimensions. Thus, our competitive layer is represented as a two-dimensional array of neurons as shown in Figure 2.
4.2.2. Parameters of the Neuron Struct
The neurons in the two-dimensional array mentioned above have a data structure with the following properties:
List of links to adjacent neurons There are 6 adjacent neurons for any given neuron, for they are placed on a hexagonal grid. Array indices for adjacent neurons can be computed using a fairly simple function for a plane SOM, but are unable to compute in some cases for a spherical SOM, namely at the edges of the net. Therefore we store the addresses of the adjacent neurons as a list for later reference.
Weight vector The weight vector is stored as a one-dimensional array.
Euclid distance This property temporarily holds the computed distance between a neuron and a given input data point.

Figure 2. Description of spherical surface on a two-dimensional array.
Trained flag This flag is turned on if a neuron has already been trained with a given input data point to avoid redundant exposures.
Validity flag This flag is turned off if a neuron is not a valid part of competitive layer, in which case it should be excluded when computing distances and winner neurons.
Three-dimensional coordinates These coordinates are used to draw a three dimensional sphere.
4.2.3. Distance and Winner Neuron Computation
Because the neurons are placed in a two-dimensional array, distances and BMUs can be computed by simply scanning this array, as with a plane SOM. They are filtered before computation, however, according to their validity flags.
4.2.4. Choosing Neurons to Be Trained
A SOM is trained by choosing neurons that are within a certain distance from a given point and adjusting them. In order to choose these neurons, we construct a search tree that has a certain depth by recurring into the link of adjacent neurons, thus representing a range of neurons to be trained. The tree is then traversed breadth first so that neurons closer to the origin (winner neuron) are trained first. Below are the details of this process.
First, the winner neuron is trained. Then, we retrieve all adjacent neurons of the winner neurons and train them. At the same time, a list of neurons trained is created as shown in Figure 3. Then, we train the neurons that are adjacent to the ones in the list, effectively moving outward (these neurons are also stored in a new list).
The trained flag is checked before each neuron is trained so that one neuron does not get trained multiple times from one input data point. Thus we recursively train neighboring neurons of the winner without duplications or omissions.
4.2.5. Displaying the Competitive Layer
It is often effective to colorize each neuron when displaying the competitive layer as a visualization. We mapped the weight vector of each of the neurons into the HSV color space to colorize the neurons. The Saturation and Value were fixed to the maximum, while the Hue was determined from the weight vector. The Hue can take any value in range [0, 360]. We used values in range [0, 280] so that large values show up as red and small values show up as purple.
5. Execution of Spherical SOM with GPUs
5.1. Hardware Acceleration
The SOM algorithm is known to have high parallelism for its distance computation, winning neuron computation, and training in each cycle of its execution. Therefore we can expect fast execution using recent technologies such as multi-core processors and other hardware suitable for parallel computation.
High-speed executions of SOMs have been much debated since the well-known research by Carpenter, et al. on hardware acceleration in 1987 [7]. COKOS (Coprocessor for Kohonen’s Self-organizing map), a SIMD processor specifically for SOMs, was developed by Speckman in 1992 [8]. This processor had 8 parallel processing cores and was designed to make the most out of SOM’s parallelism.
There have also been attempts to construct a SOM accelerator on FPGA by Porrmann and Tamukoh [9]. Porrman constructed a piece of hardware that executes SOMs with 6 processing modules connected to each other in a ring topology. Tamukoh implemented a piece of hardware that chooses a winner neuron by computing all distances at once and evaluated it using a vector dimension number of 128 and a map size of 16 by 16. He also used Manhattan distances instead of Euclidian distances to simplify computations and reduce circuit size. Processing modules and data memories existed for each weight vector, and the way these modules connected with each other replicated the competitive layer. Thus the modules were able to compute in a massively parallel manner. This piece of hardware could compute a SOM in constant time regardless of map size, and was proved to be 350 times faster than an Intel Xeon 2.80 GHz CPU.
In 2009 Shitara attempted a high-speed execution of SOM using a GPU. He used an NVIDIA GeForce GTX- 280, which was 150 times faster than a CPU [10]. This high parallelism holds for spherical SOMs as well. In this study we use GPUs for which development environments

Figure 3. Choosing neurons to be trained.
have been published (and are actively researched) to implement a spherical SOM and discuss the performance based on execution with GPUs and conventional CPUs.
5.2. Parallelizing the SOM Algorithm
Let us discuss which parts of the SOM algorithm may be parallelized.
The most costly process in the algorithm is the computations of Euclidian distances between input vectors and neurons’ weight vectors. These computations are independent from each other so they may be parallelized.
Selecting the winner neuron, the process in which a neuron with the minimum Euclidian distance is determined, may also be parallelized. Sequential comparison would be O(n), but this can be reduced to O(log(n)) by employing a tournament-style comparison with binary trees.
Training of the neurons may also be parallelized. In this process, computations against weight vectors for all neurons within a certain distance from the winner neuron are to be performed for a given input data point. These adjustments are independent of each other and may be parallelized.
5.3. Evaluation
Based on the above facts, we implemented a spherical SOM program that is executable with a GPU, and evaluated its execution speed.
C++, along with CUDA, NVIDIA’s GPU development environment, was used in implementation. The GPUs we used are 1) GeForce 8400GS, 2) GeForce GTX280, and 3) Tesla C1060. Table 1 shows their technical specifications.
We also implemented a spherical SOM program for CPU to compare. The CPU we used was 4) Opteron 1210 HE.
SOM settings were as follows:
• 5-dimensional neurons
• 50,000 (20 × 502) weight vectors
• [0,9] neighborhood range
5.3.1. Results
Table 2 shows execution times for each GPU and CPU.