1. INTRODUCTION
Crude oil constitutes a major source of pollution in our environment [1]. And its accidents nowadays have become a common phenomenon and caused ecological and social catastrophes [2], and reported to influence the biodiversity, distribution and pollution of Microorganisms in an environment [3].
In the last two decades, there have been increased public concerns on the adverse effect of oil exploration on the environment. The toxic effects of crude oil and refined petroleum oils on plants, animals, humans and the environment are devastating [4]. Oil pollution persistence and its transport in water, subsoil and groundwater aquifers are monitored to predict impacts, assess the impacts and audit such effects with a view to mitigate the impacts. Environmental monitoring of petroleum hydrocarbons pollution ranges from specific methods, such as the use of radio-active labeled compounds to general methods including quantifying gross contamination and evaluating the extent of change caused in the environment by the presence of the pollutant [5].
Microbial degradation is the major and ultimate natural mechanism by which one can cleanup hydrocarbon pollutants from the environment [6,7].
Biodegradation is the metabolic utilization of organic compounds by microorganisms which require oxygen, nutrients, water and specified physico-chemical conditions such as moderate temperature and pH [8].
Microbial populations increase rapidly in the water after an oil spill. Oil spilt on the ground can rapidly disappear completely under optimal conditions often within a year or two as a result of microbial oxidation of the hydrocarbons. Consequently it may be useful to employ the natural microflora in cleaning-up of oil spills [9].
Several hundred species of yeasts, moulds, bacteria and actinomycetes are now known to possess the capacity to oxidize hydrocarbons. Since biodegradation is a natural process, it has many advantages with few ecological side effects: it is relatively inexpensive with low energy requirements and it can be carried out without elaborate equipment [10]. In addition, it is cheaper than other remediation technologies and believed to be noninvasive and relatively cost-effective [11].
Bacteria have the ability to exploit carbon compounds in oil as a necessary source of energy [10], and have large capability to adapt to the environment that live in it by using its own special enzymatic system, so it is widespread compared to other microorganisms [12]. The aim of this study was to explore efficiency of Bacillus thuringiensis in biodegradation of light crude oil.
2. MATERIAL & METHODS
2.1. Culture Media
2.1.1. Mineral Salt Medium (MSM)
This media was used for growth of bacterial isolate that used in the experiment which consists of: 1 g (NaCl), 1 g (KH2PO4), 1 g (Na2HPO4), 0.5 g (NH4NO3), 0.5 g ((NH4)2SO4), 0.2 g (MgSO4∙7H2O), 0.02 g (CaCl2∙2H2O), 0.002 g (FeCl3), 0.002 g (MnSO4∙2H2O), which all dissolve in 1 Liter of distilled water and pH has been justify to 7 and sterilizes by autoclave [13].
2.1.2. Modified Mineral Salt Medium (MMSM)
This media was used to encourage oil emulsion process which leads to oil degradation. This media consist of: 4 g (NH4NO3), 4 g (KH2PO4), 0.2 g (MgSO4∙7H2O), 0.01 g (CaCl2∙2H2O), which all dissolves in 1 Liter of distilled water and pH has been justify to 7 and sterilizes by autoclave [14].
2.1.3. Lactose-Broth
This media was used to activate the preserved bacterial isolate, this media consist from: 10 g (Tryptone), 5 g (yeast extract), 5 g (NaCl), 1 g (Glucose), which all dissolve in 1 Liter of distilled water and pH has been justify to 7.2. then, 25 ml from this solution transferred to containers and sterilizes by autoclave [15].
2.1.4. Nutrient Agar
Nutrient agar prepared by dissolving 9.2 g of Agar powder in 400 ml of Distilled Water as mentioned in kit instruction of Oxoid Company (England). This medium was used for growth of bacterial isolate to count their numbers.
2.2. Study the Morphological Changes of Crude Oil Degradation by Bacterial Isolate
Mineral Salt Medium (MSM) was used for growing of B. thuringiensis bacteria, 50 ml form Mineral Salt Medium was transfer to conical flasks (3 replicates). Then the MSN were cooled and sterilized, after that 2 ml from crude oil and 2 ml from bacteria broth were added. Flasks were incubated at 30˚C for 27 day [16].
2.3. Viable Count of Bacteria Isolate
A series of dilutions were done by dissolving 0.1 g of bacteria in 10 ml of liquid MSM. 0.1 ml from the 7th dilution spreads on plates of nutrient agar Medium, and the plates were incubated in 30˚C. Viable count was calculated after the inoculation directly (Zero Time) up to 4 week. The viable count was calculated for growing bacteria colonies by Italian Colony Counter [17].
2.4. Estimate the Amount of Bioemulsion Produced by Bacteria
Three bottles contain 50 ml of sterile liquid MMSM had been inoculated by bacterial isolate, then 2 ml of crude oil were added, the bottles were incubated in vibration incubator in 30˚C at 150 r/min for 27 days. The emulsion amount was estimated by expelled samples by centrifuge at 12,000 r/min for 30 minutes. Then the clear layer (represented the amulsion) was pulled by syringe carefully. pH was adjusted to acidic value (2) and left for 24 hours at 4˚C. The filtrate layer was transferred to a separating funnel and then added chloroform and methanol solvent. Finally, to get rid of the solvent, the extract was put in the oven at 45˚C - 40˚C until it dries. The dried extract was weighted which represents the amount of emulsion expressed in grams [18].
2.5. Measuring the Quantitative Loss Percentage of Crude Oil Using Bacteria
Measuring the Quantitative loss percentage of crude oil using bacteria by non-specialized chemical methods and tested the ability of bacterial isolate for the degradation of crude oil and was done by two ways.
2.5.1. Account of Biomass
After bacterial isolate were grow on the activation medium for 24 hours, 50 ml of sterile liquid MSM was inoculated by bacterial isolate in conical flask, then added 2 ml of crude oil (Three replicate). Flasks were placed in vibration incubator at 150 r/min in 30˚C for 27 days. The bacterial broth put in a special tube and then expelled by Centrifuge at 12,000 r/min for 30 minutes to precipitate the bacterial cells. Cells were extracted with a mixture of acetone and hexane 1:3 to rid it of hydrocarbons that adherent with it. Cells were dried at 105˚C for 24 hours in a thermal oven. Finally, dry weight method was used to estimate the weight of the biomass [19].
2.5.2. Account the Quantitative Loss of Crude Oil as a Result of the Growth of Bacterial Isolate
The pure bacterial isolate was inoculated in conical flasks contain 50 mil sterile liquid MSM, and 2 ml of crude oil were added, Then incubated in the vibration incubator at 30˚C in 150 r/min for 27 days and by three replicates, then expelled by Centrifuge at 12,000 r/min for 30 minutes to precipitate the bacterial cells. After sedimentation of cells the filtrate collected with hexane extract and then adding a few drops of hydrochloric acid to reach pH to 2. Hydrocarbons extracted by adding Chloromethane solvent after putting the filtrate in a separating funnel to separate the water layer from hydrocarbons. The Hydrocarbons extract was filtrated through filter paper that contains anhydrous sodium sulfate (Na2SO4) as dryer factor, the Hydrocarbons extract was collected in a clean beaker, and then the solvent was evaporated in room temperature using a rotary evaporator. The Hydrocarbons extract was weighted to calculate the remaining weight of hydrocarbons, this procedure was returned by using a control sample, and the percentage rate of this degradation was determined by the equation below:

PRDCO = Percentage Rate of Degradation of Crude Oil [20].
2.6. Detection the Degradation of Crude Oil by Specialized Chemical Methods Using a Technology of Gas-Chromatography
Bacterial isolate was inoculated in 50 ml of sterile liquid MSM in conical flask after it’s grown on the activation Medium for 24 hours, and then 2 ml of crude oil were added (three replications). Flasks was incubated in the vibration incubator at 150 r/min at 30˚C for 27 days, the bacterial broth transfer to a special tube, then expelled by Centrifuge at 12,000 r/min for 30 minutes to precipitate the cells, Then the filtrate was dissolved with Hexane 1:1. After that, the filtrate was dried by using a rotary evaporator at 80˚C. The final sample (filtrate) which represents the remainder of the hydrocarbons was tested by Gas-Chromatography device that equipped by the Shimadzu Japanese Company, Model 2014 using Chromatography separation column type (Capillary column CPSIL5-CB). The resulted areas were calculated by equation bellow to estimate the amount of hydrocarbons remaining after microbial degradation [18]:

Remaining Percentage of hydrocarbons = (sample area)/ (total area) × 100
3. RESULTS & DISCUSSION
3.1. Morphological Changes in Crude Oil Degraded by Bacterial Isolate
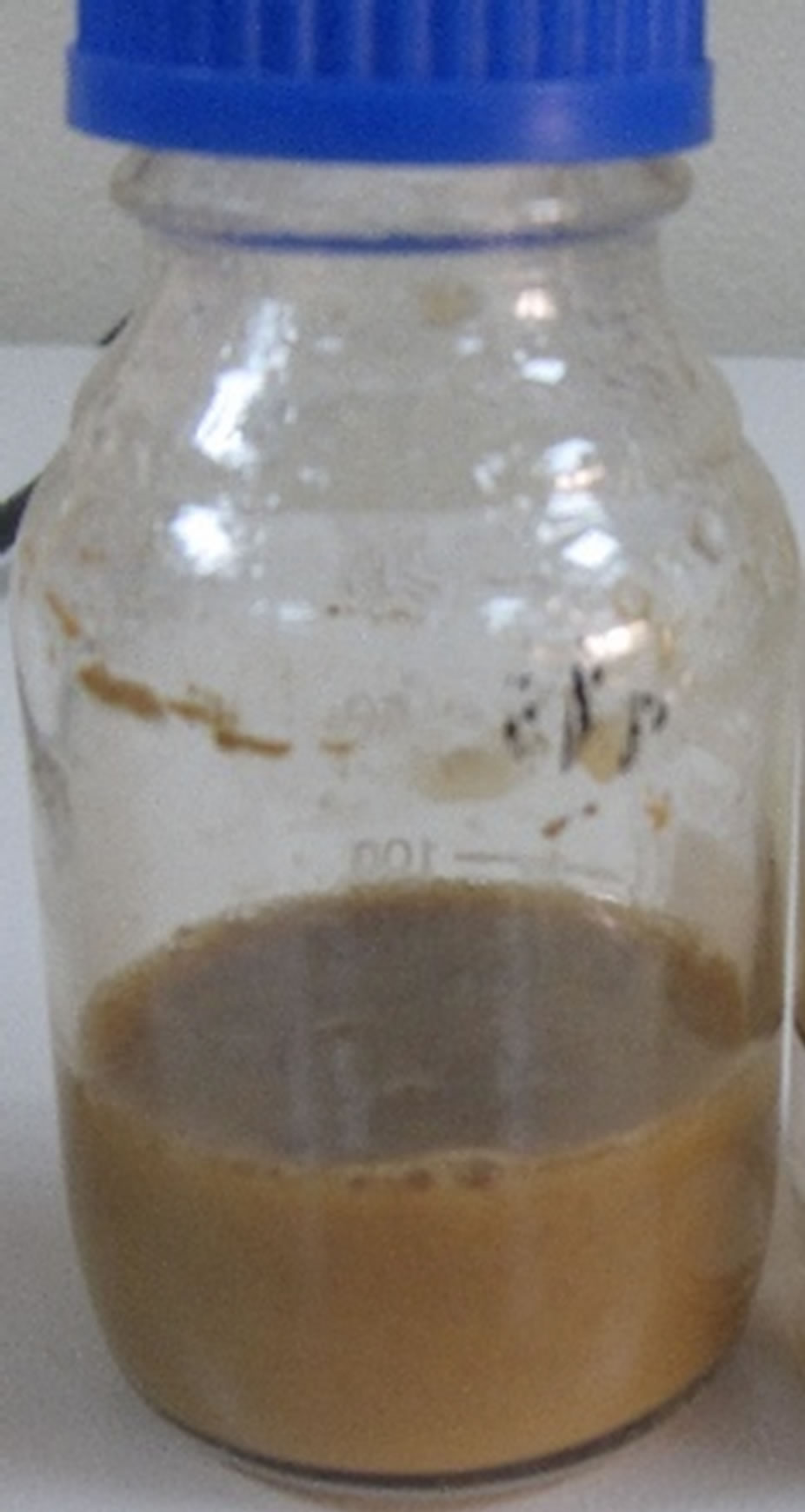
The results shows superficial changes in the layer of light crude oil due to the growth of bacterial after incubation period for 27 days at 30˚C, the layer was shifted in mass of crude oil to gelatinous mass and loss its strength and emulsification, compared to a control sample (Plate 1). This proves the ability of the bacteria in degradation of crude oil as a result of production of bioemulsions [21-23].
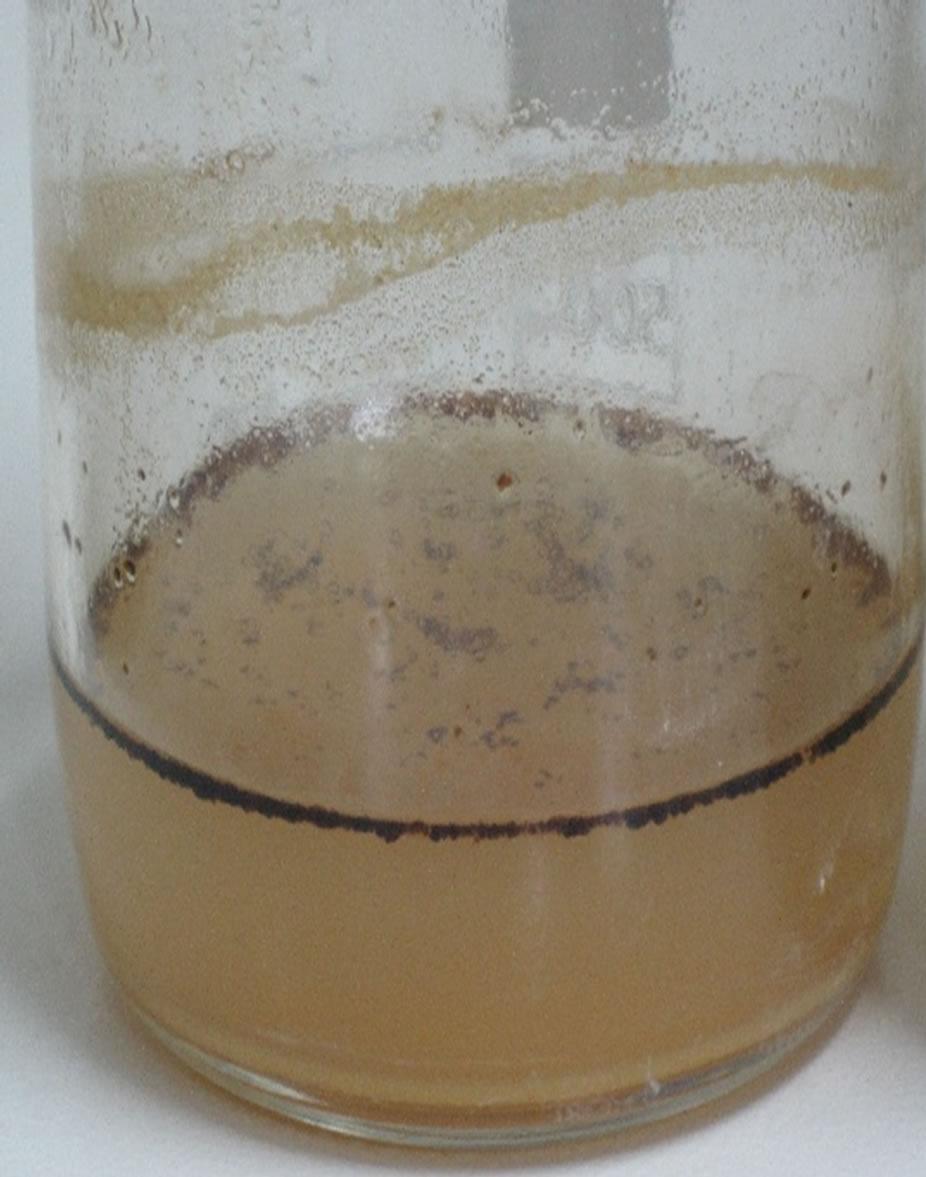
The conglomerate of crude oil in form of gelatinous mass is called pseudo-solubilization, and it occurs as a result of the accumulation of Emulsifying agents which scatter the oil in the form of small droplets that spread and form emulsion (Plate 2). According to Kosaric [16] these materials are characterized by: biodegradability, generally low toxicity, changing surface active phenomena, such as lowering of surface and interfacial tensions, wetting and penetrating actions, spreading, hydro-phylicity and hydrophobicity actions, microbial growth enhancement, metal sequestration and anti-microbial action.
3.2. Viable Count of Bacteria That Degraded Crude Oil
Increasing numbers of bacteria with the progress of time is considering one of the methods that used to determine the extent of their ability to adaptation and degradation of crude oil compounds [8]. The total number of
 (a)
(a)  (b)
(b)
Plate 1. Morphological changes of crude oil degraded by B. thuringiensis that incubated at 30˚C for 27 days on liquid MSM. (a) Control sample; (b) Treated sample with B. thuringiensis.
 (a)
(a)  (b)
(b)
Plate 2. Morphological changes of crude oil degraded by B. thuringiensis that incubated at 30˚C for 27 days on liquid MSM. (a) Control sample; (b) Scattered of crude oil in the form of small droplets that spread and form emulsion by action of B. thuringiensis.
viable count has been continually increased over the four weeks where it was impossible to determine their numbers after the third week due to the high density, so the viable count was increased with incubation time (Table 1).
This may be due to the availability of oil compounds that were subjected to bacterial decomposition, which are considered a food source to the bacteria [24,25], because the crude oil gives bacteria a source of carbon and energy [10] and also to adapt the bacteria to an environment that contaminated with crude oil [26], and to the presence of bacterial enzymes [27]. These enzymes enable microorganisms to combust the hydrocarbons. The genetic machinery needed to make these oil-degrading enzymes is most commonly found in bacteria (although many fungi and some other organisms can also degrade oil). The bacterial cell then harnesses the released energy by degrading the compounds to support its own life processes. The situation is exactly analogous to the way our bodies break down the chemical energy in food to provide the energy and raw materials for maintenance, growth, and repair of our tissues [28].
There are many different varieties of hydrocarbons, and over millions of years, bacteria has evolved catalytic machines (enzymes) that are specific for particular degradation reactions. Some of the simpler compounds can be degraded by a very wide variety of bacteria, but the ability to degrade other compounds (hydrocarbons, for example) is found in fewer species. No one bacterium can make all the different enzymes, instead; each kind of bacterium specializes in a few hydrocarbons as preferred food sources [29].
Most microbial oil degradation occurs by aerobic respiration, in other words, the oil-degrading microbes breathe oxygen and burn oil hydrocarbons just as humans breathe oxygen and burn food for energy. In the absence of oxygen, microbes have other mechanisms to degrade hydrocarbons for energy. Biodegradation of oil constituents without oxygen (i.e., under anoxic conditions) is much slower but anoxic processes may be relevant to the longterm restoration efforts (e.g., in oil contaminated salt marsh environments). From the human point of view, the microbes is degrading or breaking down the oil, which results in cleaning up the environment that has been contaminated by the spill. From the microbial point of view, they metabolizing or consuming the oil to provide the energy and materials that needed to live and grow [1].
3.3. Bioemulsion Produced by B. thuringiensis
Bioemulsifiers are composed of polysaccharides, protein, lipopolysaccarides or complex mixtures of these polymers. These microbial emulsifiers are important secondary metabolism produced by microbes [30]. Bioemulsifiers have effectiveness impressive at the level of surface tension, leading to emulsification of hydrocarbon compounds and make them available to the microorganisms [31].
In this study, the amounts of emulsion produced by B. thuringiensis reached 2.3 g/l. while, in other studies the amounts of emulsion produced by Bacillus subtilis reached 2 g/l [32] and 1.47 g/l [33]. So, the B. thuringiensis was efficient producers of Bioemulsifiers in a rich hydrocarbons medium. Biosurfactant secreted by bacteria are more effective than chemical surfactants in enhancing the solubility and biodegradation of petroleum hydrocarbons [34,35]. Production of Bioemulsifiers was related to the utilization of available hydrophobic substrates by the producing microbes from their natural habitat, presumably by increasing the surface area of substrates and increasing their solubility phenomenon [36-39].
3.4. Quantitative Losing Percentage of Crude Oil Using B. thuringiensis Bacteria
As mentioned in the material and methods it has been measure the Quantitative losing percentage of crude oil by B. thuringiensis bacteria using non-specialized chemical methods and tested the ability of bacterial isolate on the dismantling of crude oil in two ways.
3.4.1. Account of Biomass
Al-Khazali [22] Proved when conducting studies on B. subtilis if the biomass increases for more than 3 g/l This indicates that the bacteria are highly efficient in a Oil degradation. In this study the total biomass of bacterial isolate reached 5 g/l, when grown on culture medium that supplied with crude oil as a source of carbon and energy. So, B. thuringiensis can be highly efficient in the process of degradation the oil. The results show a shift in color of medium to turbid and this is evidence on the growth of B. thuringiensis that used in this experiment and their ability to consume the medium components, compared with a control sample [40]. Microbes break oil down into simple carbon compounds that are used to make the sugars, fats, and proteins needed for growth and energy production, ultimately the byproducts become carbon dioxide and water. The simple carbon compounds are incorporated into new cellular constituents, in other words, they are used to make more microbes. So, the carbon has been used to produce additional biomass [1].
3.4.2. Account the Quantitative Loss of Crude Oil as a Result of the Growth of B. thuringiensis
The results show that the B. thuringiensis have the ability to degraded 80% of the weight of the crude oil used in the experiment, while Bacillus subtilis in the study of [29,41] have ability to degraded 65%, 64% of the weight of the crude oil, respectively. This may be due to the adaptation of B. thuringiensis to the environment and production of emulsion materials [42] and to the presence of bacterial enzymes [43], also to the availability of optimal temperature, which raise the efficiency of bacteria in the degradation of the crude oil components [44].
3.5. Detection of the Degradation of Crude Oil by Specialized Chemical Methods Using a Technology of Gas-Chromatography
By using gas-Chromatography device, it appears in the control sample presence of 43 chemicals material (Table 2). While In contrast, the sample that biodegraded by Bacillus thuringiensis reveled extensive biodegradation in 27 days, the visual examination of gas-Chromatography shows the disappearance of a number of beams where the number of chemical materials have been decreased to 26 (Table 3), also decreased in peak area for some material, compared with a control sample (Figures 1 and 2). This clearly indicated that B. thuringiensis is a good hydrocarbons decomposers. This method is a specialized chemical method that which inferred the type of decaying compound [45] and clarify the decomposition that happens in oil compounds [46,47].

Table 1. Viable count for 7th dilution of B. thuringiensis that degraded crude oil incubated in 30˚C.


Table 2. Numbers of Chemical composition of crude oils in control sample.

Table 3. Numbers of Chemical composition of crude oils in the sample that injected with Bacillus thuringiensis.

Figure 1. Showed the appearance of Chemical material in control sample that appeared by using gas-Chromatography.
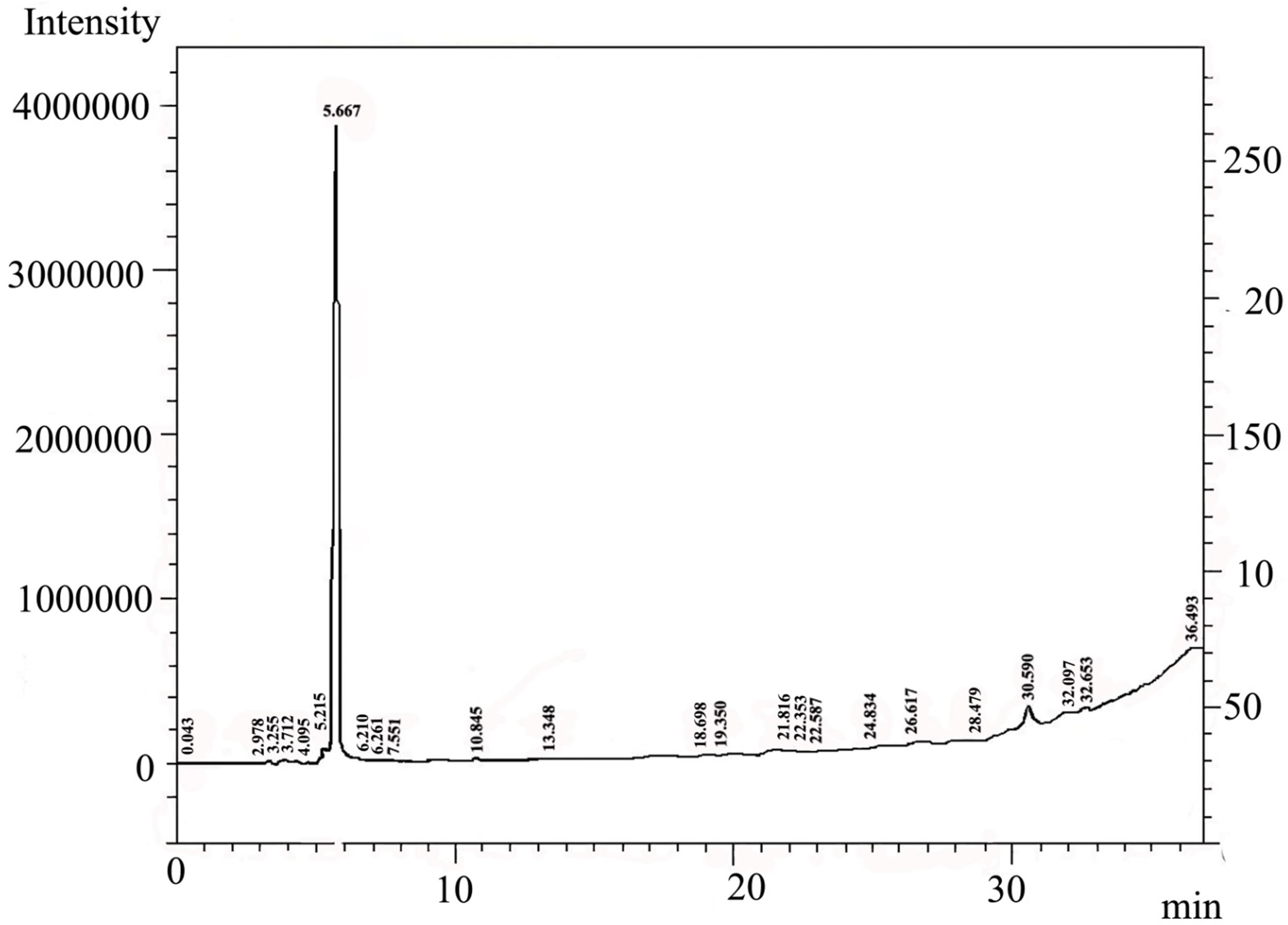
Figure 2. Showed the appearance of Chemical material in the sample that injected with Bacillus thuringiensis by using gas-Chromatography.
4. CONCLUSION
In the study, changes that occurred in the morphological, viable count and biomass, of the oil-degrading bacteria were examined. Bacillus thuringiensis bacteria were used under the optimal conditions in order to determine their oil-degrading ability and they underwent different tests in the laboratory environment in medium rich with hydrocarbonate. The investigation revealed that bacteria had the ability to degrade crude oil, by utilizing oil products used as substrates. Bacteria exhibited best development processes and had the highest levels of oil-degradation, because the set-up of this experiment emulated a natural environment. It is a significant investigation that may light the way for bioremediation studies and provides information that allows the employment of Bacillus thuringiensis for bioremediation in environments polluted with crude oil. However, further studies need to be carried out to know the composition of Bioemulsion produced by Bacillus thuringiensis, and its positive and negative effects on aquatic ecosystem.
5. ACKNOWLEDGEMENTS
The authors are thankful to ministry of science and technology and Baghdad University to Permit to carry out the study.