American Journal of Plant Sciences
Vol.4 No.1(2013), Article ID:27573,6 pages DOI:10.4236/ajps.2013.41001
Applying Molecular Breeding to Improve Soybean Rust Resistance in Vietnamese Elite Soybean
![]()
1Department of Molecular Biology, Agricultural Genetics Institute, Hanoi, Vietnam; 2Institute of Agricultural Science for Southern Vietnam, Ho Chi Minh City, Vietnam; 3Graduate School for International Development and Cooperation, Hiroshima University, Hiroshima, Japan.
Email: *tdxuan@hiroshima-u.ac.jp
Received October 17th, 2012; revised November 26th, 2012; accepted December 14th, 2012
Keywords: Rust Resistant Gene; Soybean; Marker Assisted Selection; Molecular Marker
ABSTRACT
Soybean production in Vietnam has recently been threatened by the widespread dissemination of soybean rust (SBR) caused by the fungus Phakopsora pachyrhizi. Application of molecular breeding is considered as a feasible method to improve soybean rust resistance and minimize the adverse effects from overuse fungicides in this country. In this study, we have successfully applied molecular markers in a backcross breeding program to introgress the Rpp5 gene of SBR resistance into HL203, an elite Vietnamese soybean variety, from two donor lines of DT2000 and Stuart 99084B-28. The plants in BC4F1 generation had maximum contribution from the recurrent parents and retained SBR resistance gene.
1. Introduction
Soybean (Glycine max (L) Merr.) is one of the most important crops in the world, accounting for 48% of the world market in oil crops, and is widely used for animal feed for oil and protein production [1]. In Vietnam, soybean is an important crop and is utilized as a good source of meal food and high-quality protein. However, soybean production is severely reduced because of soybean rust (SBR) which is caused by Phakopsora pachyrhizi Syd. & P. Syd. P. pachyrhizi reproduces asexually on soybean and all other known leguminous host plants through the production of dikaryotic urediniospores [1]. The disease is disseminated through urediniospores carried by the wind and can develop rapidly, causing loss of poliar area and a severe reduction in worldwide soybean grain yield. In some areas where the disease is common, the yield losses can be up to 80% [2]. Hence, SBR is considered a formidable threat to world soybean production [1]. This pathogen has first reported to detect in Asia and widely spread many areas in the world [2,3]. Chemical spray containing fungicides is the only effective method to control the disease but increases production costs and is not an environmental friendly approach and appearing some P. pachyrhizi increased tolerance to the certain fungicides [4]. Therefore, introduction of the resistant varieties is the most effective measure to control this disease, the use of resistant variety is the most feasible solution because it is simple to manage and friendly to the environment. To establish suitable varieties, plant breeders should optimize the plant genotype by choosing the most promising resistance genes and combinations to ensure durability of resistance [5]. Scientists worldwide have screened many soybean accessions for resistance or tolerance to soybean rust [6,7] and identified five different loci carrying dominant alleles: Rpp1 identified in PI 200492 [8], Rpp2 from (PI 230970) [9], Rpp3 (PI 230970) [10], Rpp4 (PI 459025) [11] and Rpp5 (PI 200487 and PI 471904) [12]. Another recent research has identified recessive genes controlling SBR resistance [13]. Besides single-gene resistance, partial growth and sporulation may be controlled by genes with minor effect [3,14].
Introgression of desired genes by the traditional protocols is lengthy and complicated. Molecular markers are considered valuable tools for crop improvement, due to their usefulness in characterizing and manipulating genetic loci responsible for monogenic and polygenic traits [15]. The use of molecular markers can be useful tool to monitor the transfer of alleles of interest [16,17], and can speed up the development of soybean varieties carrying single or multiple resistance genes. The microsatellite markers have been used in soybean for mapping specific genes that determine agronomic traits, and also identified QTLs of economic importance, involving in grain yield and genetic resistance to pests and diseases, which are characteristics of complex inheritance [18].
Soybean has a reasonably dense molecular-marker linkage map [19] and the association of marker to the known genes has been pursued by many groups. Molecular mapping of SBR-resistance genes in soybean has previously been reported. Brogin et al. (2004) [20] identified single Sequence Repeat (SSR) markers linked to rust resistance present on the variety FT-2 in the linkage group (LG)-C2 of the previous soybean consensus map reported by Cregan et al. (1999) [21]. However, the locus could not be identified in the study. An SBR resistance gene from the variety Hyuuga was mapped at 3cM interval on LG-C2 between Satt134 and Satt460 [22]. Hyten (2007) [23] recently mapped the Rpp3 locus at the same interval as the report of Monteros et al. (2007) [22]. The Rpp1 locus has been mapped to a 1 cM interval on LG-G between Sct_187 and Sat-064 LG-G [23].
There is a lack of information on gene pyramiding by applying maker assisted selection (MAS) for soybean rust in Vietnam. Therefore in this work, the newly developed lines that were obtained and exhibited SBR resistance and retained the yield and grain quality traits of HL203. This study represents a successful example of the use of molecular markers, in foreground and background selection to introgress the genes of interest into a premium soybean variety.
2. Materials and Methods
2.1. Plant Materials and Breeding Scheme
Resistant analysis was performed in two backcrossing populations obtained from two donors, DT2000 and Stuart 99084B-28. The recurrent parent was a SBR susceptible variety, HL203, which is an elite soybean variety, and is widely grown in the South Vietnam because of its logging and drought resistance, and high yield (2.2 - 2.5 tons/ha) (data not shown). The breeding scheme is shown in Figure 1.
2.2. Inoculation, Phenotype and Molecular Marker Analysis
The isolated using in this study was obtained by collecting spore from naturally infected greenhouse plants of the susceptible variety cultured in M1 medium with a total volume of 1 L including 10 g glucose, 1 g·K2HPO4, 5 g peptone, 0.5 MgSO4·7H20, 20 g agar and adjust sterile distilled water of 1L. Fungal colonies were transferred from a master plate to two pre-prepared M1 medium plate and incubated at room temperature for 36 - 48 h. The experiment soybean lines were designed by randomized complete block design (RCBD) with at least three replications; two leaves of each plant per replication were infected.

Figure 1. Breeding schemes were produced for developing backcrossing populations containing SBR resistance gene with genetic background of HL203.
After 8 - 10 days, evaluation of the affected level was recorded following by standard protocol of IRRI. Performance of rust disease on leaves was grouped using NTSYS pc software version.
Healthy leaf tissue was collected from the parents and backcrossing plants. Tissue was frozen in liquid nitrogen, freeze-dried, and ground to a fine powder using a modified CTAB protocol [24]. The DNA was precipitated with isopropanol and treated with Rnase A. DNA concentration and integrity was estimated by spectrophotometer analysis and gel electrophoresis, respectively. Simple sequence repeat (SSR) molecular markers were selected based on the reported genomic location of the known Rpp genes. SSR primer sequences were obtained from soyBase [25].
For SSR analysis, 30 µg of DNA was used as template in a 10 µl reaction containing buffer (100 mM Tris-HCL, 500 mM KCL), 1.5 mM·MgCL2, 32.5 µM of each dNTP, 0.2 µM of each primer, and 1 U of Taq DNA polymerase. The cycling consisted of 5 min at 94˚C; 35 cycles of 1 min at 94˚C, 1 min at 50˚C, 1 min at 72˚C; followed by 7 min at 72˚C. The amplified fragments were separated by electrophoresis in 3% agarose, stained with ethidium bromide, and visualized under UV light.
3. Results
3.1. Introduction of SBR Resistance Gene into HL203 Background
Rpp5 locus located in N linkage group between franking markers Sat_275 and Sat_280 shown in Figure 2. The DT2000 and Stuart 99084B-28 soybean varieties (donor of the SBR resistance Rpp5 gene) were crossed to HL203 with the former as the male parent. The F1 plants were confirmed for their heterozygosis for the “R” gene linked markers and were backcrossed using HL203 as a female parent.
The resulting BC1F1 lines were first checked for presence of the marker linked to Rpp5 resistance allele in a heterozygous condition. All of the plants that were heterozygous for Rpp5 were backcrossed to HL203 to generate BC2F1 plants (HL203 as female parent) and the process was continued up to the BC4F1 stage. A representative example of genotyping for background selection is provided in Figure 1. At the BC4F1 generation, the plant having maximum contribution from the recurrent parent and containing “R” gene of SBR resistance was selfed to obtain BC4F2 lines that were screened using the “R” gene linked marker to identify plants that were homozygous for “R” gene of Rpp5 resistance Figures 3 and 4.
3.2. Evaluation of SBR Resistance to Backcrossing Generation
The backcrossing lines were evaluated for their resistance to SBR under glass house condition using several different isolates of SBR. As compared to HL203, the leaves of the backcrossing lines containing Rpp5 gene exhibited very small lesion lengths indicating the high level of resistance to SBR (data not shown).
The lines containing Rpp5 exhibited good level of resistance. Mudge et al. (1997) [27] reported that microsatellite franking Satt038 and Satt130 allele rhg1 and Sat- 168 and Satt 309 delimiting rgh1. Recently, Garcia et al.

Figure 2. Genetic linkage map constructed with SSR markers. (a) Represents a fragment of the consensus soybean linkage group N; (b) Illustrates the map derived from the F2 population CD 208 × PI 200456; (c) Shows the map derived from the F2 population CD 208 × PI 471904; (d) Illustrates the map derived from the F2 population CD 208 × PI 200526. The lines indicate the corresponding position of the markers between the four maps. The arrows show the position of the soybean rust resistance genes. The genetic distances (cM) are shown on the left [12,26].

Figure 3. PCR detection of Rpp5 gene in representative in BC4F1 plant. Marker used was Sat_275 (M: 100 bp ladder, 1: HL203 recurrent parent, 2: DT2000 donor, 3 - 13: BC4F1 plants).
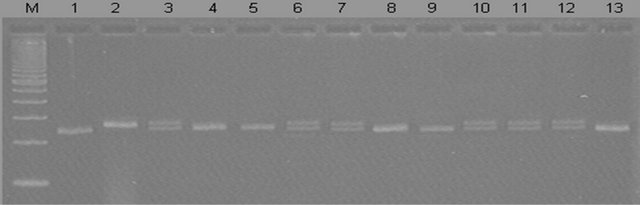
Figure 4. Genotyping for detection of Rpp5 gene in representative in BC4F1 plant using marker Sat_275 (M: 100 bp ladder, 1: HL203 recurrent parent, 2: Stuart 99084B-28 donor, 3 - 13: BC4F1 plants).
(2008) [12] mapped the locus Rpp5 in three different populations of soybean (PI 200456, PI 471904 and PI 200526) and obtained six markers have linkage to this locus. Also, two new microsatellite markers showed potentially involved in resistance of soybean to soybean rust and found the link to Rpp5 gene and are present on the linkage group N of soybean. The efficiency of selection was recorded for all markers linked to gen Rpp5, and the combination of the markers Sat_275 + Sat_280 was 100% [28]. Application of molecular markers is using as valuable tools for crop improvement, due to its usefulness in characterizing and manipulating genetic loci responsible for monogenic and polygenic traits. Marker assisted selection (MAS) used in this study has successfully introgressed Rpp5 resistance gene into HL203. MAS to the introgression of genes from the donor (the donor genotype) to another (the recurrent genotype) via a backcrossing breeding program clearly illustrates the powerful advantages of the use of molecular marker for indirect selection of trait interest which would otherwise be difficult to select by the standard procedures [15].
4. Discussion
HL203 is a regional soybean variety which is popularly grown due to its high yield, grain quality and high adaptability in soybean cultivation regions of South Vietnam. In the present study, Rpp5 resistance gene was introgressed into HL203 with the objective of developing SBR resistant lines and retaining the yield and quality characteristics of HL203.
Sporedical reports have documented about the distribution of soybean rust in Vietnam before 2005; however, recently Pham et al. (2010) [7] screened various selected soybean genotypes for resistance to soybean rust in Vietnam. It indicated some potentially useful sources of resistance to soybean rust that may be valuable to soybean breeders. Nevertheless, this work is the first report in Vietnam which has demonstrated that marker assisted backcrossing can be gainfully employed for adding new genes into popular and elite soybean varieties that have been grown by farmers over the years on account of their unique characteristics. It can be expected that the availability of the soybean genome sequence will facilitate the development of many more markers for transfer of traits of agronomic value, with greater precision, into comercially important soybean varieties.
5. Acknowledgements
The authors express sincere thanks to the grant from Ministry of Agriculture and Rural Development of Vietnam to this research.
REFERENCES
- M. R. Miles, R. D. Frederick and G. Hartman, “Soybean Rust: Is the US Soybean Crop at Risk?” APSnet Feature, American Phytopathological Society, 2003. http://www. apsnet. org/online/feature/rust/
- L. Li, J. R. Smith, J. D. Ray and R. D. Frederick, “Identification of a New Soybean Rust Resistance Gene in PI 567102B,” Theorical and Applied Genetics, Vol. 125, No. 1, 2012, pp. 133-142. doi:10.1007/s00122-012-1821-y
- Y. Ono, P. Buritica and J. F. Hennen, “Delimitation of Phakopsora, Physopella cerotelium and Their Species on Leguminosae,” Mycological Research, Vol. 96, No. 10, 1992, pp. 825-850. doi:10.1016/S0953-7562(09)81029-0
- C. Godoy, “Changes in Performance of Soybean Rust Fungicides over Years and New Management Strategies Adopted in Brazil,” National Soybean Rust Symposium, New Orleans, 9-11 December 2009.
- A. Palloix, V. Ayme and B. Moury, “Durability of Plant Major Resistance Genes to Pathogens Depends on the Genetic Background, Experimental Evidence and Consequences for Breeding Strategies,” New Phytologist, Vol. 183, No. 1, 2009, pp. 190-199. doi:10.1111/j.1469-8137.2009.02827.x
- M. R. Miles, W. Morel, J. D. Ray, J. R. Smith, R. D. Frederick and G. L. Hartman, “Adult Plant Evaluation of Soybean Accessions for Resistance to Phakopsora pachyrhizi in the Field and Greenhouse in Paraguay,” Plant Disease, Vol. 92, No. 1, 2008, pp. 96-105. doi:10.1094/PDIS-92-1-0096
- T. A. Pham, C. B. Hill, M. R. Miles, B. T. Nguyen, T. T. Vu, T. D. Vuong, T. T. V. Toai, H. T. Nguyen and G. L. Hartman, “Evaluation of Soybean for Resistance to Soybean Rust in Vietnam,” Field Crop Research, Vol. 117, No. 1, 2010, pp. 131-138. doi:10.1016/j.fcr.2010.02.011
- R. J. Mclean and D. E. Byth, “Inheritance Resistance to Rust (Phakopsora pachyrhizi) in Soybeans,” Australia Journal Agricultural Research, Vol. 31, No. 5, 1980, pp. 951-956. doi:10.1071/AR9800951
- K. R. Bromfield and J. S. Melching, “Sources of Specific Resistance to Soybean Rust,” Phytopatology, Vol. 71, 1982, p. 706 (Abstract).
- K. R. Bromfield and E. E. Hartwig, “Resistance to Soybean Rust and Mode of Inheritance,” Crop Science, Vol. 20, No. 2, 1980, pp. 254-255. doi:10.2135/cropsci1980.0011183X002000020026
- E. E. Hartwig, “Identification of a 4th Major Gene Conferring Resistance to Soybean Rust,” Crop Science, Vol. 26, No. 6, 1986, pp. 1135-1136. doi:10.2135/cropsci1986.0011183X002600060010
- A. Garcia, E. S. Calvo, R. A. de Souza Kiihl, A. Harada, D. M. Hiromoto and L. G. Vieira, “Molecular Mapping of Soybean Rust (Phakopsora pachyrhizi) Resistance Genes: Discovery of a Novel Locus and Alleles,” Theorical and Applied Genetics, Vol. 117, No. 4, 2008, pp. 545-553. doi:10.1007/s00122-008-0798-z
- E. S. Calvo, R. A. S. Kiihl, A. Garcia, A. Harada and D. M. Hiromoto, “Two Major Recessive Soybean Genes Conferring Soybean Rust Resistance” Crop Science, Vol. 48, No. 4, 2008, pp. 1350-1354. doi:10.2135/cropsci2007.10.0589
- G. L. Hartman, M. R. Miles and R. D. Frederick, “Breeding for Resistance to Soybean Rust,” Plant Diseases, Vol. 89, No. 6, 2005, pp. 664-666. doi:10.1094/PD-89-0664
- A. M. Torres, “Application of Molecular Markers for Breeding Disease Resistant Varieties in Crop Plants,” In: S. M. Jain and D. S. Brar, Eds., Molecular Techniques in Crop Improvement, Springer Science and Business Media B.V, Dordrecht, 2010.
- S. D. Tanskley, “Molecular Markers in Plant Breeding,” Plant Molecular Biology Reporter, Vol. 1, No. 1, 1983, pp. 3-8. doi:10.1007/BF02680255
- S. D. Tanskley and S. R. Mc Couch, “See Banks and Molecular Maps: Unlocking Genetic Potential from the Wild,” Science, Vol. 277, No. 5329, 1997, pp. 1063-1066. doi:10.1126/science.277.5329.1063
- J. Yuan, V. N. Njiti, K. Meksen, M. J. Iqbal, K. Triwitayakorn, M. A. Kassem, G. T. Davis, M. E. Schimdt and D. A. Lightfood, “Quantitative Trait Loci in Two Soybean Recombinant Inbred Line Populations Segregating for Yield and Disease Resistance,” Crop Science, Vol. 42, No. 1, 2002, pp. 271-277. doi:10.2135/cropsci2002.2710
- Q. J. Song, L. F. Marek, R. C. Shoemaker, K. G. Lark, V. C. Concibido, X. Delannay, J. E. Specht and P. B. Cregan, “A New Interagted Genetic Linkage Map of the Soybean,” Theorical and Applied Genetics, Vol. 109, No. 1, 2004, pp. 122-128. doi:10.1007/s00122-004-1602-3
- R. L. Brogin, C. A. A. Arias, N. A. Vello, J. F. F. Toledo, A. E. Pipolo, L. L. Catelli and S. R. R. Marin, “Molecular Mapping of a Gene Conferring Resistance to Soybean Rust,” Poster Presented at the VII World Soybean Research Conference, Foz do Iguassu, 29 February-5 March 2004.
- P. B. Cregan, T. A. L. Jarvik, A. L. Bush, R. C. Shoemaker, K. G. Lark, A. L. Kahler, et al., “An Integrated Genetic Linkage Map of the Soybean Genome,” Crop Science, Vol. 39, No. 5, 1999, pp. 1464-1490. doi:10.2135/cropsci1999.3951464x
- M. J. Monteros, A. M. Missaoui, D. V. Phillips, D. R. Walker and H. R. Boerma, “Mapping and Confirmation of the ‘Hyuuga’ Red-Brown Lesion Resistance Gene for Asian Soybean Rust,” Crop Science, Vol. 47, No. 8, 2007, pp. 829-836. doi:10.2135/cropsci06.07.0462
- D. L. Hyten, G. L. Hartman, R. L. Nelson, R. D. Frederick, V. C. Concibido, J. M. Narvel and P. B. Cregan, “Map Location of the Rpp1 Locus That Confers Resistance to Soybean Rust in Soybean,” Crop Science, Vol. 47, No. 2, 2007, pp. 837-840. doi:10.2135/cropsci2006.07.0484
- P. Keim, T. C. Olson and R. C. Shoemaker, “A Rapid Protocol for Isolating Soybean DNA,” Soybean Genetic Newsletter, Vol. 15, 1988, pp. 150-152.
- Soybase, 2012. http://soybase.org/resources/ssr.php
- Q. J. Song, L. F. Marek, R. C. Shoemaker, K. G. Lark, V. C. Concibido, X. Delannay, J. E. Specht and P. B. Cregan, “A New Integrated Geneticlinkage Map of the Soybean,” Theorical and Applied Genetics, Vol. 109, No. 1, 2004, pp. 122-128. doi:10.1007/s00122-004-1602-3
- J. Mudge, P. B. Cregan, J. P. Kenworthy, W. J. Kenworthy, J. H. Orf and N. D. Young, “Two Microsatellite Markers That Flank the Major Soybean Cyst Nematode Resistance Locus,” Crop Science, Vol. 37, No. 5, 1997, pp. 1611-1615. doi:10.2135/cropsci1997.0011183X003700050034x
- T. G. S. Morceli, S. H. Unêda Trevisoli Jr., A. A Morceli, R. A. S. Kiihl, E. S. Calvo, A. O. Di Mauro and A. Garcia, “Identification and Validation of Microsatellite Markers Linked to the Rpp5 Gene Conferring Resistance to Asian Soybean Rust,” Pesquisa Agropecuária Brasileira, Vol. 43, No. 11, 2008, pp. 1533-1541. doi:10.1590/S0100-204X2008001100011
NOTES
*Corresponding author.

