Attractor-Based Simultaneous Design of the Minimum Set of Control Nodes and Controllers in Boolean Networks ()
Received 20 June 2016; accepted 15 August 2016; published 18 August 2016

1. Introduction
Modeling, analysis, and control of gene regulatory networks are one of the fundamental problems in the field of systems biology. Control of gene regulatory networks is closely related to therapeutic interventions, which are realized by radiation, chemotherapy, and so on. Hence, developing control theory of gene regulatory networks is important for gene therapy technologies (see, e.g., [1] ) in the future. When we discuss the control problem of gene regulatory networks, we frequently assume that some of genes can be arbitrarily controlled. In this paper, such genes are called a control node. Since in mathematical models of gene regulatory networks, control nodes are not necessarily specified, the problem of finding a set of control nodes based on control specifications is important as a kind of inverse problems. Needless to say, it is desirable that the number of control nodes is small.
In this paper, we study simultaneous design of a minimum set of control nodes and controllers in a Boolean networks (BN). A BN [2] is one of the simplified models such as Bayesian networks and piecewise affine systems [3] , and is widely used in theoretical analysis of gene regulatory networks (see, e.g., [4] - [19] ). In the BN, dynamics such as interactions between genes are expressed by Boolean functions, that is, gene expression is expressed by a binary value (0 or 1). As a generalized version of BNs, a probabilistic BN has been proposed in [20] . In the simultaneous design problem studied in this paper, control specifications on attractors are imposed. Attractors represent cell types or states of cells [21] . In particular, we focus on a singleton attractor, which is also called a fixed point. By solving this problem, we can obtain a gene regulatory network that has desired attractors and has no undesired attractors. Furthermore, we can obtain time evolution (i.e., a Boolean function) of a control node, i.e., a controller by solving this problem. Thus, a solution of the simultaneous design problem provides a helpful suggestion to e.g., design of therapeutic interventions.
As a related problem, the conventional controllability problem of BNs has been widely studied (see, e.g., [7] [12] [13] [17] [18] [22] [23] ). However, the problem of finding a minimum set of control nodes has not been considered. In [24] , using piecewise affine systems, the problem of choosing a control node has been studied. However, the minimization problem of the number of control nodes has not been considered.
In order to solve the simultaneous design problem, a matrix-based representation of BNs [10] is utilized. Using this representation, this problem can be rewritten as an integer linear programming problem. We may use a matrix-based representation using the semi-tensor product (STP) method [7] [25] . Since manipulation of matrices using the STP is not needed in this paper, we utilize a simple matrix-based representation using truth tables. The effectiveness of the proposed method is presented by an example on a WNT5A network, which is related to melanoma.
Notation: For the finite set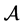 , let
, let 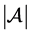 denote the number of elements in
denote the number of elements in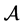 . For the n-dimensional vector
. For the n-dimensional vector 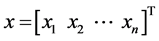 and the index set
and the index set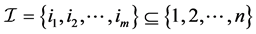 , define
, define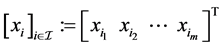 . For two matrices A and B, let
. For two matrices A and B, let  denote the Kronecker product of A and B. In addition, for q vectors
denote the Kronecker product of A and B. In addition, for q vectors 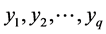 and the index set
and the index set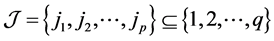 , define
, define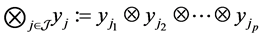 . For example, for q two-dimensional vectors
. For example, for q two-dimensional vectors 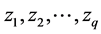 and
and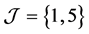 , we can obtain
, we can obtain
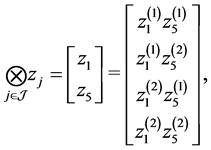
where 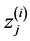 is the i-th element of
is the i-th element of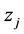 . Finally, let
. Finally, let ![]() denote the
denote the ![]() matrix whose elements are all one.
matrix whose elements are all one.
2. Boolean Networks
Consider the following Boolean network (BN):
![]() (1)
(1)
where ![]() is the state (e.g., the expression of genes), and
is the state (e.g., the expression of genes), and ![]() is the dis-
is the dis-
crete time. The set ![]() is a given index set, and expresses the adjacency relation of nodes corre-
is a given index set, and expresses the adjacency relation of nodes corre-
sponding to genes. The function ![]() is a given Boolean function consisting of logical operators such as AND (
is a given Boolean function consisting of logical operators such as AND (![]() ), OR (
), OR (![]() ), and NOT (
), and NOT (![]() ). If
). If ![]() holds, then
holds, then ![]() is uniquely determined as 0 or 1.
is uniquely determined as 0 or 1.
Next, the notion of attractors is defined as follows.
Definition 1 The state ![]() is called a singleton attractor if
is called a singleton attractor if ![]() holds.
holds.
Definition 2 The set of states ![]() is called a periodic attractor if
is called a periodic attractor if![]() ,
, ![]() and
and ![]() hold.
hold.
We present a simple example.
Example 1 Consider the following BN:
![]() (2)
(2)
In this BN, ![]() ,
, ![]() , and
, and ![]() hold. Figure 1 shows the state transition diagram. From Figure 1, we see that there exist two singleton attractors and one periodic attractor with
hold. Figure 1 shows the state transition diagram. From Figure 1, we see that there exist two singleton attractors and one periodic attractor with![]() . ,
. ,
In this paper, we focus on only a set of singleton attractors. Hereafter, let ![]() denote a set of singleton attractors for the BN (1).
denote a set of singleton attractors for the BN (1).
3. Problem Formulation
The states ![]() are decomposed to two groups
are decomposed to two groups![]() ,
, ![]() and
and![]() ,
, ![]() , where
, where ![]() and
and ![]() are index sets satisfying
are index sets satisfying ![]() and
and![]() . Then, defining
. Then, defining![]() ,
, ![]() , the BN (1) can be rewritten as
, the BN (1) can be rewritten as
![]() (3)
(3)
where![]() ,
, ![]() ,
, ![]() , and
, and ![]() are an index set. For each
are an index set. For each![]() , the relations
, the relations ![]() and
and ![]() hold. The Boolean function
hold. The Boolean function![]() ,
, ![]() is given in advance. The Boolean function
is given in advance. The Boolean function![]() ,
, ![]() is not given, and must be found. In other words, we consider that some of Boolean functions in the BN (1) are replaced with new Boolean functions. In the case of
is not given, and must be found. In other words, we consider that some of Boolean functions in the BN (1) are replaced with new Boolean functions. In the case of![]() , (3) is the same as (1). Furthermore, in the BN (3),
, (3) is the same as (1). Furthermore, in the BN (3), ![]() corresponds to the control node (i.e., the control input). The Boolean function
corresponds to the control node (i.e., the control input). The Boolean function![]() ,
, ![]() can be regarded as the controller.
can be regarded as the controller.
Then, consider the following problem.
Problem 1 For the BN (3), suppose that Boolean functions ![]() are given. Suppose also that the set of desired singleton attractors and the set of undesired singleton attractors are given by
are given. Suppose also that the set of desired singleton attractors and the set of undesired singleton attractors are given by
![]() (4)
(4)
and
![]() (5)
(5)
![]()
Figure 1. State transition diagram, where the number assigned to each node implies![]() .
.
respectively. Then, find a minimum set of control nodes and a controller![]() ,
, ![]() satisfying the following two conditions:
satisfying the following two conditions:
![]() (6)
(6)
and
![]() (7)
(7)
If this problem is infeasible, then there exists no controller that realizes specifications on singleton attractors. This fact implies that the BN (3) is uncontrollable from the viewpoint of singleton attractors, and the change of the graph structure ![]() is required. On the other hand, as a solution,
is required. On the other hand, as a solution, ![]() may be obtained. This solution implies that a given BN has desired singleton attractors and has no undesired singleton attractors. That is, control nodes are not necessary. Thus, Problem 1 can be regarded as a kind of the controllability problems.
may be obtained. This solution implies that a given BN has desired singleton attractors and has no undesired singleton attractors. That is, control nodes are not necessary. Thus, Problem 1 can be regarded as a kind of the controllability problems.
If the set ![]() (i.e.,
(i.e.,![]() ) is given, Problem 1 has been already solved in [10] . Problem 1 is more general than the problem studied in [10] . By solving Problem 1, we can obtain both the minimum set of control nodes and the controller. To the best of our knowledge, such problem has not been studied so far.
) is given, Problem 1 has been already solved in [10] . Problem 1 is more general than the problem studied in [10] . By solving Problem 1, we can obtain both the minimum set of control nodes and the controller. To the best of our knowledge, such problem has not been studied so far.
4. Matrix-Based Representation of Boolean Networks
In order to solve Problem 1, we use a matrix-based representation of BNs [10] .
In this representation, one state ![]() in the BN (1) is expressed by two binary variables
in the BN (1) is expressed by two binary variables ![]() and
and![]() . The binary variable
. The binary variable ![]() is defined by
is defined by![]() . The binary variable
. The binary variable ![]() is defined by
is defined by![]() . Using
. Using ![]() and
and![]() , define
, define
![]()
Then, the matrix-based representation for ![]() is given by
is given by
![]() (8)
(8)
where ![]() and
and![]() . The matrix
. The matrix ![]() can be derived from truth tables. We present a simple example.
can be derived from truth tables. We present a simple example.
Example 2 Consider the BN (2) again. Then, we can obtain the truth table for each![]() . See Table 1. From these truth tables, we can obtain the following matrix-based representation:
. See Table 1. From these truth tables, we can obtain the following matrix-based representation:
![]()
![]()
![]()
where each element of![]() ,
, ![]() is given by a binary value (0 or 1), and a sum of all elements in each column of
is given by a binary value (0 or 1), and a sum of all elements in each column of ![]() is equal to 1. The second row in
is equal to 1. The second row in![]() ,
, ![]() corresponds to the column in
corresponds to the column in ![]() in truth tables. ,
in truth tables. ,
See [10] for further details of deriving the matrix![]() .
.
The matrix-based representation is useful for considering Problem 1. Using the matrix-based representation, the problem of finding a Boolean function is reduced to the problem of finding![]() . The latter problem is relatively easier than the former problem, and can be solved by using some computational tools.
. The latter problem is relatively easier than the former problem, and can be solved by using some computational tools.
Remark 1 Recently, there have been many results about a matrix-based representation of Boolean functions using the semi-tensor product (STP) of matrices (see, e.g., [7] [25] ). Of course, the matrix-based representation using the STP method may be utilized. In this paper, we focus on singleton attractors (i.e., the steady state), and time sequences of the state are not computed. Hence, manipulation of matrices using the STP is not needed, and the simple matrix-based representation using truth tables is utilized.
5. Solution Method for Problem 1
Based on the matrix-based representation of BNs explained in the previous section, we consider a solution method for Problem 1.
As preparations, the notation is defined. If ![]() in (8) includes decision variables, then
in (8) includes decision variables, then ![]() is denoted by
is denoted by![]() . Let
. Let ![]() and
and ![]() denote the j-th element of
denote the j-th element of ![]() and
and ![]() in
in ![]() of (4) and
of (4) and ![]() of (5), respectively. De- fine
of (5), respectively. De- fine
![]()
Since only singleton attractors are focused in Problem 1, Problem 1 is equivalently transformed into the following problem.
Problem 2 Find matrices ![]() and binary variables
and binary variables ![]() minimizing the cost function
minimizing the cost function ![]() subject to the following two conditions. (i) For each
subject to the following two conditions. (i) For each![]() , the following constraints corresponding to (6) are satisfied:
, the following constraints corresponding to (6) are satisfied:
![]()
(ii) For each![]() , at least one of the following constraints is satisfied:
, at least one of the following constraints is satisfied:
![]()
In addition, each element of ![]() takes a binary value, and a sum of each column of these matrices is equal to 1. ,
takes a binary value, and a sum of each column of these matrices is equal to 1. ,
In this problem, the condition (i) guarantees that![]() ,
, ![]() , that is,
, that is, ![]() ,
, ![]() are included in the set of singleton attractors. The condition (ii) guarantees that
are included in the set of singleton attractors. The condition (ii) guarantees that![]() ,
, ![]() are not a singleton attractor. If
are not a singleton attractor. If![]() ,
, ![]() are obtained as a solution, then this solution implies
are obtained as a solution, then this solution implies![]() , that is, the controller is not needed. If
, that is, the controller is not needed. If![]() ,
, ![]() are obtained as a solution, then this solution implies
are obtained as a solution, then this solution implies![]() , that is, all Boolean functions are changed.
, that is, all Boolean functions are changed.
Problem 2 can be rewritten as an integer linear programming (ILP) problem, and can be solved by a suitable solver. An SMT (Satisfiability Modulo Theories) solver such as the Yices SMT Solver [26] can also be applied to Problem 2.
Here, using a simple example, we explain a method to rewrite Problem 2 as an ILP problem. Consider the BN in Example 2 of Section 4. Then, the constraints appeared in Problem 2 are given by
![]() (9)
(9)
![]() (10)
(10)
![]() (11)
(11)
and
![]() (12)
(12)
![]() (13)
(13)
![]()
where ![]() is a binary decision variable. From (9)-(11), we can obtain
is a binary decision variable. From (9)-(11), we can obtain
![]() (14)
(14)
![]() (15)
(15)
![]() (16)
(16)
From (12)-(14), we can obtain
![]() (17)
(17)
![]() (18)
(18)
![]() (19)
(19)
From the condition (ii) in Problem 2, at least one of (17)-(19) must be satisfied. Noting this fact, from (17)-(19) we can obtain
![]() (20)
(20)
![]() (21)
(21)
![]() (22)
(22)
where![]() ,
, ![]() ,
, ![]() are additional binary variables satisfying the two constraint conditions
are additional binary variables satisfying the two constraint conditions
![]() (23)
(23)
Under these conditions, at least one of![]() ,
, ![]() takes either −1 or 1. Thus, Problem 2 for the BN in Example 2 can be rewritten as the following problem.
takes either −1 or 1. Thus, Problem 2 for the BN in Example 2 can be rewritten as the following problem.
![]()
subject to (14)-(16), (20)-(22), and (23).
Finally, the product of binary variables such as ![]() can be linearized by using the following lemma [27] .
can be linearized by using the following lemma [27] .
Lemma 1 Suppose that binary variables![]() ,
, ![]() are given, where
are given, where ![]() is a given index set. Then
is a given index set. Then
![]() is equivalent to the following linear inequalities
is equivalent to the following linear inequalities
![]()
where ![]() is the cardinality of
is the cardinality of![]() .
.
Using this lemma, Problem 2 for the BN in Example 2 can be rewritten as an ILP problem. From the above, we see that Problem 2 for a general BN (3) can be rewritten as an ILP problem. In general, an ILP problem is NP-hard, but several free/commercial solvers were developed. Hence, we can solve the ILP problem obtained.
6. Example on a WNT5A Network
In this section, as an example, we consider the gene regulatory network with the gene WNT5A, which is related to melanoma. A BN model is given by
![]()
![]()
![]()
![]()
![]()
![]()
![]()
where the concentration level (high or low) of the gene WNT5A is denoted by![]() , the concentration level of the gene pirin by
, the concentration level of the gene pirin by![]() , the concentration level of the gene S100P by
, the concentration level of the gene S100P by![]() , the concentration level of the gene RET1 by
, the concentration level of the gene RET1 by![]() , the concentration level of the gene MART1 by
, the concentration level of the gene MART1 by![]() , the concentration level of the gene HADHB by
, the concentration level of the gene HADHB by![]() , and the concentration level of the gene STC2 by
, and the concentration level of the gene STC2 by![]() . See [28] for further details.
. See [28] for further details.
For this BN model, consider solving Problem 1. We assume that![]() , and
, and ![]() are given by
are given by
![]()
![]()
respectively. This setting is artificially given.
First, we derive a matrix-based representation. Then, we can obtain the following matrix-based representation:
![]()
![]()
![]()
![]()
![]()
![]()
![]()
Next, we explain the solution for Problem 2. By solving this problem, we can obtain ![]() and
and![]() ,
,![]() . That is, the minimum value of
. That is, the minimum value of ![]() is obtained by 1, and the sets
is obtained by 1, and the sets ![]() and
and ![]() are obtained by
are obtained by ![]() and
and![]() , respectively. This implies that only the Boolean function
, respectively. This implies that only the Boolean function ![]() must be replaced with other Boolean function. As the solution, we can obtain
must be replaced with other Boolean function. As the solution, we can obtain
![]()
Finally, we observe the set of singleton attractors. For the BN model obtained, the set of singleton attractors is given by
![]()
From this set, we see that specifications on singleton attractors (![]() of (6) and
of (6) and ![]() of (7)) are satisfied.
of (7)) are satisfied.
7. Conclusions
For Boolean networks, we studied the problem of finding both a minimum set of control nodes and a controller, where specifications on singleton attractors are imposed. In biological networks, control nodes (control input) are not necessarily given. Hence, this problem is important in both systems biology and synthetic biology. Using a matrix-based representation of BNs, this problem can be rewritten as an integer linear programming problem. The proposed method is effective as one of the fundamental methods for control theory of gene regulatory networks.
There are several open problems. In this paper, we focused on only the set of singleton attractors. An extension of the proposed method to the case of the set of periodic attractors is important, but it will be a challenging topic. Utilizing a probabilistic BN as a model of gene regulatory networks is also important. Then, a discrete probability distribution of attractors must be considered. Finally, in order to clarify the effectiveness of the proposed method, it is important to consider applying the proposed method to several biological systems.
Acknowledgements
This research was partly supported by JSPS Grant-in-Aid for Scientific Research (C) 26420412.