Modeling and Simulation for High Energy Sub-Nuclear Interactions Using Evolutionary Computation Technique ()
Received 19 November 2015; accepted 10 January 2016; published 13 January 2016

1. Introduction
Evolutionary computation refers to a class of algorithms that utilize simulated evolution to some degree as a means to solve a variety of problems, from numerical optimization to symbolic logic. By simulated evolution, we mean that the algorithms have the ability to evolve a population of potential solutions such that weaker solutions are removed and replaced with incrementally stronger (better) solutions. In other words, the algorithms follow the principle of natural selection. Each of the algorithms has some amount of biological plausibility, and is based on evolution or the simulation of natural systems [1] - [8] .
In 1990s, John Koza [9] [10] introduced the subfield called Genetic Programming. This is considered a subfield because it fundamentally relies on the core genetic algorithm created by Holland [11] , and differs in the underlying representation of the solutions to be evolved. Instead of using bit-strings (as with genetic algorithms) or real-values (as is the case for evolutionary programming [1] - [7] or evolutionary strategies [1] - [7] , genetic programming relies on S-expressions (program trees) as the encoding scheme.
Hadron-nucleus (h-A) interactions have been considered as a corner stone in high energy physics because of its theoretical and practical interesting features and also it is an intermediate state between hadron-hadron (h-h) and nucleus-nucleus (N-N) interactions. So, there are a lot of models that concern the study of the hadron structure [12] - [16] and the interactions between hadrons and nuclei such as the three-fireball model [17] , quark models [18] - [20] , fragmentation model [21] - [23] and many more.
In our previous works [24] - [27] , our group studied the applications of artificial intelligence and the evolutionary computation techniques such as neural network, adaptive fuzzy inference system, genetic programming, genetic algorithm, hybrid technique model and many others to solve many complex (nonlinear) problems in high energy physics and showed best fitting with the corresponding experimental data in comparison with the conventional techniques.
The study of hadron-nucleus interaction at high and ultrahigh energy has been a subject of great interest to high energy physicist because the nuclei provide a number of unique physics opportunities which are not available in elementary particle collisions [28] .
In this article, Genetic programming (GP) model has been used to discover a function that computes the rapidity distribution of created (total charged, positive and negative) pions for p−-Ar and p−-Xe collisions at 200 GeV/c [29] - [31] and charged particles for p-pb collision at 5.02 TeV [32] . The seven discovered functions produced by GP model show an excellent matching when they have been compared to the corresponding experimental data [29] - [32] . This article is organized as follows; Section 2 gives the definition and the outlines to the basics of the GP technique. Section 3 reviews the implementation of GP. Finally, the results and conclusions are provided in Sections 4 and 5 respectively.
2. GP Outlines
GP is defined as the biologically-inspired evolution of computer programs that solve a predefined task. For this reason, GP is nothing more than a genetic algorithm applied to the problem program evolution. Early GP systems utilized LISP (the original functional programming language, 1958) S-expressions (as shown in Figure 1), but more recently, linear GP systems have been used to evolve instruction sequences to solve user-defined programming tasks [2] .
Since genetic programming manipulates programs by applying genetic operators (reproduction, crossover and mutation), a programming language should permit a computer program to be manipulated as data and the newly created data to be executed as a program. For these reasons, LISP was chosen as the main language for GP.
Evolving complete programs with GP is computationally very expensive, and the results have been limited, but GP does have a place in the evolution of program fragments. For example, the evolution of individual functions that have very specific inputs and outputs and whose behavior can be easily defined for fitness evaluation by GP. To evolve a function, the desired output must be easily measurable in order to understand the fitness landscape of the function in order to incrementally evolve it [1] - [7] .
Consider the example shown in Figure 1. The population consists of two members, A and B. Using the crossover operator, a portion of A is grafted onto B; resulting in a new expression C. GP also utilizes the mutation operator as a way of extending the population to the search space.
![]()
Figure 1. Using the crossover operator to create a new S-expression.
3. GP Implementation
The GP uses the same fundamental flow as the traditional genetic algorithm. The population of potential solutions is initialized randomly and then their fitness computed (through a simulation of executed instructions with the stack). Selection of members that can propagate into the next generation can then occur through fitness proportionate selection. With this method, the higher fit the individual, the higher the probability that they will be selected for recombination in the next generation. Evolutionary algorithms borrow concepts from Darwinian natural selection as a means to evolve solutions to problems, choosing from more fit individuals to propagate to future generations [1] - [7] .
The chromosome, or program to be evolved, is made up of genes, or individual instructions. The chromosome can also be of different lengths, assigned at creation, and then inherited during the evolution.
All methods of evolutionary computation (and then GP) work as follows: create a population of individuals, evaluate their fitness, generate a new population by applying genetic operators (Cross-over, mutation and reproduction), and repeat this process a number of times as shown in Figure 2.
4. Results and Discussion
We have performed the GP modeling of the inclusive reaction,

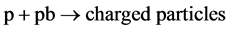 (1)
(1)
using the experimental data [29] - [32] at 100 and 200 GeV/c and have done so many runs to select the best runs of the GP program, the first runs are for simulating the rapidity distribution 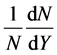 of negative pions for
of negative pions for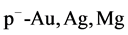 collisions at 100 GeV/c. They were configured to have the lab momentum
collisions at 100 GeV/c. They were configured to have the lab momentum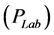 , mass number (A) and the number of particles per unit solid angle (Y) as inputs and the output is the corresponding rapidity distribution
, mass number (A) and the number of particles per unit solid angle (Y) as inputs and the output is the corresponding rapidity distribution 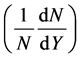 of negative pions at the given momentum as shown in Figure 3.
of negative pions at the given momentum as shown in Figure 3.
The second ones are for simulating the rapidity distribution 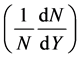 of positive pions for
of positive pions for  collisions at 100 GeV/c as an output and the inputs are the same as in Figure 3.
collisions at 100 GeV/c as an output and the inputs are the same as in Figure 3.
The third ones are for simulating the rapidity distribution 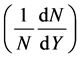 of positive pions for
of positive pions for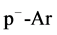 ,
, 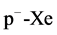
![]()
Figure 3. A block diagram of the GP technique.
collisions at 200 GeV/c as an output and the inputs are the same as in Figure 3.
The fourth ones are for simulating the rapidity distribution ![]() of negative pions for
of negative pions for![]() ,
, ![]() collisions at 200 GeV/c as an output and the inputs are the same as in Figure 3.
collisions at 200 GeV/c as an output and the inputs are the same as in Figure 3.
The fifth ones are for simulating the rapidity distribution ![]() of charged pions for
of charged pions for![]() ,
, ![]() collisions at 200 GeV/c as an output and the inputs are the same as in Figure 3.
collisions at 200 GeV/c as an output and the inputs are the same as in Figure 3.
The last ones are for simulating the rapidity distribution ![]() of charged particles for p-pb collisions at 5.02 TeV as an output and the inputs are the same as in Figure 3.
of charged particles for p-pb collisions at 5.02 TeV as an output and the inputs are the same as in Figure 3.
According to all the runs, we have obtained the corresponding tree and their equivalent discovered functions Equations (2)-(7) generated for the rapidity distribution ![]() of total charged, positive and negative pions for
of total charged, positive and negative pions for![]() collisions at 100, 200 GeV/c and 5.02 TeV.
collisions at 100, 200 GeV/c and 5.02 TeV.
The output, the rapidity distribution![]() , as a function of the inputs
, as a function of the inputs ![]() is given as follows:
is given as follows:
For negative pions at 100 GeV/c,
![]() (2)
(2)
For positive pions at 100 GeV/c,
![]() (3)
(3)
For positive pions at 200 GeV/c,
![]() (4)
(4)
For negative pions at 200 GeV/c,
![]() (5)
(5)
For charged pions at 200 GeV/c,
![]() (6)
(6)
For charged particles at 5.02 TeV,
![]() (7)
(7)
For more details about F, E,H, U, etc., see Appendices 1-6.
The comparison between the pions rapidity distribution ![]() computed by employing our discovered functions Equations (2)-(7) and the corresponding experimental data [29] - [31] are represented in Figure 4 for negative pions for
computed by employing our discovered functions Equations (2)-(7) and the corresponding experimental data [29] - [31] are represented in Figure 4 for negative pions for ![]() collisions at 100 GeV/c, Figure 5 for positive pions for
collisions at 100 GeV/c, Figure 5 for positive pions for ![]()
collisions at 100 GeV/c, Figure 6 for positive pions for ![]() (the GP model cannot describe the data when the axial value is near 0 because the noisy behavior of data around the axial),
(the GP model cannot describe the data when the axial value is near 0 because the noisy behavior of data around the axial), ![]() collisions at 200 GeV/c. Figure 7 for negative pions for
collisions at 200 GeV/c. Figure 7 for negative pions for![]() ,
, ![]() collisions at 200 GeV/c, Figure 8 for charged pions for
collisions at 200 GeV/c, Figure 8 for charged pions for![]() ,
, ![]() collisions at 200 GeV/c, Figure 9 for charged particles for p-pb collisions at 5.02 TeV.
collisions at 200 GeV/c, Figure 9 for charged particles for p-pb collisions at 5.02 TeV.
In order to generate the GP model we have implemented the GP steps (fitness evaluation, reproduction, crossover and mutation) that were mentioned in Section 3. Our six discovered functions are generated using the obtained control GP parameters, which are shown in Table 1.
![]()
Figure 4. The discovered rapidity distribution of negative pions ![]() for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
![]()
Figure 5. The obtained rapidity distribution of positive pions ![]() for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
![]()
Figure 6. The discovered rapidity distribution of negative pions ![]() for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
for antip-Au, antip-Ag, antip-Mg interaction at 100 GeV/c: (―) GP model, (o) experimental data.
![]()
Figure 7. GP-simulated for rapidity distribution of negative pions ![]() for antip-Ar, antip-Xe at 200 GeV/c: (―) GP model, (o) experimental data.
for antip-Ar, antip-Xe at 200 GeV/c: (―) GP model, (o) experimental data.
![]()
Table 1. Optimal parameters controlling GP program.
![]()
Figure 8. Simulated results for rapidity distribution of chared pions ![]() for antip-Ar, antip-Xe at 200 GeV/c: (―) GP model, (o) experimental data.
for antip-Ar, antip-Xe at 200 GeV/c: (―) GP model, (o) experimental data.
![]()
Figure 9. The discovered rapidity distribution of charged particles for p-Pb interaction at 5.02 TeV: (―) GP model, (o) experimental data.
The statistical error criterion of mean square error (MSE) was used to measure the deviation between the experimental (actual) and simulated values. The statistical parameter MSE has been used in this work as a performance metric [33] to compare the GP simulation with the actual observations (experimental data) and these were evaluated by using Matlab program. The smaller the values of MSE the closer the simulated values to the experimental ones. Our obtained MSE values for the seven discovered functions are given in Table 1, which show also that the performance of the GP model is clearly suitable.
5. Conclusions
GP model has been shown to be a vital method for modeling the h-A interactions. The current article presents an efficient approach for computing the rapidity distribution ![]() of charged, positive and negative pions for
of charged, positive and negative pions for ![]() and
and ![]() collisions at 200 GeV/c and charged particles for p-pb collisions at 5.02 TeV through the
collisions at 200 GeV/c and charged particles for p-pb collisions at 5.02 TeV through the
obtained discovered functions. All the discovered functions show a clear and excellent match to the experimental data.
The interaction of hadrons with atomic nuclei at high and ultrahigh energies is an issue of great importance since in-depth studies about it provide the necessary information on properties which cannot be examined by analyzing only hadron-hadron interactions.
Finally, the present work has proved that the GP approach can be employed effectively to model the h-A interactions at the given energy.
Appendices
1. Rapidity Distribution of Negative Pions for p−‑Au, p−‑Ag and p−‑Mg Interaction at 100 GeV/c
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]()
2. Rapidity Distribution of Positive Pions for p−‑Au, p−‑Ag and p−‑Mg Interaction at 100 GeV/c
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
, ![]()
3. Rapidity Distribution of Positive Pions for p−‑Xe and p−‑Ar Interaction at 200 GeV/c
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
, ![]()
4. Rapidity Distribution of Negative Pions for p−‑Ar and p−‑Xe Interaction at 200 GeV/c
![]() ,
, ![]()
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]()
5. Rapidity Distribution of Charged Pions for p−‑Xe and p−‑Ar Interaction at 200 GeV/c
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]()
6. Rapidity Distribution of Charged Particles for p-pb Interaction at 5.02 TeV
![]() ,
, ![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]() ,
,
![]() ,
, ![]() ,
, ![]()
where, ![]() is the number of particles per unit solid angle (Y),
is the number of particles per unit solid angle (Y), ![]() , lab momentum
, lab momentum![]() ,
, ![]() , mass number (A).
, mass number (A).